

You can:
| Name | CHEMBL3353464 |
|---|---|
| Molecular formula | C25H31ClN2O3 |
| IUPAC name | N-[(4-chlorophenyl)methyl]-1-[2-(3,5-dimethylphenyl)acetyl]-N-(2-methoxyethyl)-2-methylazetidine-2-carboxamide |
| Molecular weight | 442.984 |
| Hydrogen bond acceptor | 3 |
| Hydrogen bond donor | 0 |
| XlogP | 3.9 |
| Synonyms | BDBM50032314 SCHEMBL11954533 |
| Inchi Key | CCNMCOMJTSDNSG-UHFFFAOYSA-N |
| Inchi ID | InChI=1S/C25H31ClN2O3/c1-18-13-19(2)15-21(14-18)16-23(29)28-10-9-25(28,3)24(30)27(11-12-31-4)17-20-5-7-22(26)8-6-20/h5-8,13-15H,9-12,16-17H2,1-4H3 |
| PubChem CID | 68176091 |
| ChEMBL | CHEMBL3353464 |
| IUPHAR | N/A |
| BindingDB | 50032314 |
| DrugBank | N/A |
Structure | 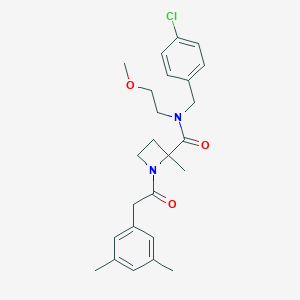 |
| Lipinski's druglikeness | This ligand satisfies Lipinski's rule of five. |
You can:
| GLASS ID | Name | UniProt | Gene | Species | Length |
|---|---|---|---|---|---|
| 443218 | Free fatty acid receptor 2 | O15552 | FFAR2 | Homo sapiens (Human) | 330 |
zhanglab![]() zhanggroup.org | +65-6601-1241 | Computing 1, 13 Computing Drive, Singapore 117417
zhanggroup.org | +65-6601-1241 | Computing 1, 13 Computing Drive, Singapore 117417