

You can:
| Name | Melatonin receptor type 1A |
|---|---|
| Species | Homo sapiens (Human) |
| Gene | MTNR1A |
| Synonym | MT1 receptor MelR Mel1a receptor Mel-1A-R |
| Disease | Insomnia Anxiety disorder Circadian rhythm sleep disorder Major depressive disorder Sleep disorders |
| Length | 350 |
| Amino acid sequence | MQGNGSALPNASQPVLRGDGARPSWLASALACVLIFTIVVDILGNLLVILSVYRNKKLRNAGNIFVVSLAVADLVVAIYPYPLVLMSIFNNGWNLGYLHCQVSGFLMGLSVIGSIFNITGIAINRYCYICHSLKYDKLYSSKNSLCYVLLIWLLTLAAVLPNLRAGTLQYDPRIYSCTFAQSVSSAYTIAVVVFHFLVPMIIVIFCYLRIWILVLQVRQRVKPDRKPKLKPQDFRNFVTMFVVFVLFAICWAPLNFIGLAVASDPASMVPRIPEWLFVASYYMAYFNSCLNAIIYGLLNQNFRKEYRRIIVSLCTARVFFVDSSNDVADRVKWKPSPLMTNNNVVKVDSV |
| UniProt | P48039 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | P48039 |
| 3D structure model | This predicted structure model is from GPCR-EXP P48039. |
| BioLiP | N/A |
| Therapeutic Target Database | T97613 |
| ChEMBL | CHEMBL1945 |
| IUPHAR | 287 |
| DrugBank | BE0000515 |
| Name | CHEMBL3580909 |
|---|---|
| Molecular formula | C14H16N2O2 |
| IUPAC name | 2-(5-methoxy-1H-indol-3-yl)-N-prop-2-enylacetamide |
| Molecular weight | 244.294 |
| Hydrogen bond acceptor | 2 |
| Hydrogen bond donor | 2 |
| XlogP | 1.9 |
| Synonyms | BDBM50096977 J3.583.566K N-Allyl-5-methoxy-1H-indole-3-acetamide AKOS029261781 |
| Inchi Key | BRKWQBQNZRMYKO-UHFFFAOYSA-N |
| Inchi ID | InChI=1S/C14H16N2O2/c1-3-6-15-14(17)7-10-9-16-13-5-4-11(18-2)8-12(10)13/h3-5,8-9,16H,1,6-7H2,2H3,(H,15,17) |
| PubChem CID | 86273229 |
| ChEMBL | CHEMBL3580909 |
| IUPHAR | N/A |
| BindingDB | 50096977 |
| DrugBank | N/A |
Structure | 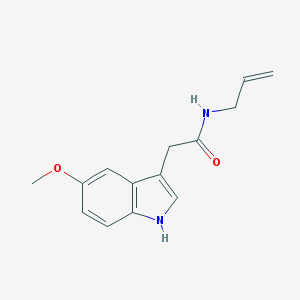 |
| Lipinski's druglikeness | This ligand satisfies Lipinski's rule of five. |
| Parameter | Value | Reference | Database source |
|---|---|---|---|
| EC50 | 525.0 nM | PMID26023814 | BindingDB,ChEMBL |
| Emax | 64.0 % | PMID26023814 | ChEMBL |
| Ki | 17.0 nM | PMID26023814 | BindingDB,ChEMBL |
zhanglab![]() zhanggroup.org | +65-6601-1241 | Computing 1, 13 Computing Drive, Singapore 117417
zhanggroup.org | +65-6601-1241 | Computing 1, 13 Computing Drive, Singapore 117417