

You can:
| Name | Metabotropic glutamate receptor 6 |
|---|---|
| Species | Rattus norvegicus (Rat) |
| Gene | Grm6 |
| Synonym | nob3 nob2 nerg1 mGluR6 mGlu6 receptor [ Show all ] |
| Disease | N/A for non-human GPCRs |
| Length | 871 |
| Amino acid sequence | MGRLPVLLLWLAWWLSQAGIACGAGSVRLAGGLTLGGLFPVHARGAAGRACGALKKEQGVHRLEAMLYALDRVNADPELLPGVRLGARLLDTCSRDTYALEQALSFVQALIRGRGDGDEASVRCPGGVPPLRSAPPERVVAVVGASASSVSIMVANVLRLFAIPQISYASTAPELSDSTRYDFFSRVVPPDSYQAQAMVDIVRALGWNYVSTLASEGNYGESGVEAFVQISREAGGVCIAQSIKIPREPKPGEFHKVIRRLMETPNARGIIIFANEDDIRRVLEATRQANLTGHFLWVGSDSWGSKISPILNLEEEAVGAITILPKRASIDGFDQYFMTRSLENNRRNIWFAEFWEENFNCKLTSSGGQSDDSTRKCTGEERIGQDSAYEQEGKVQFVIDAVYAIAHALHSMHQALCPGHTGLCPAMEPTDGRTLLHYIRAVRFNGSAGTPVMFNENGDAPGRYDIFQYQATNGSASSGGYQAVGQWAEALRLDMEVLRWSGDPHEVPPSQCSLPCGPGERKKMVKGVPCCWHCEACDGYRFQVDEFTCEACPGDMRPTPNHTGCRPTPVVRLTWSSPWAALPLLLAVLGIMATTTIMATFMRHNDTPIVRASGRELSYVLLTGIFLIYAITFLMVAEPCAAICAARRLLLGLGTTLSYSALLTKTNRIYRIFEQGKRSVTPPPFISPTSQLVITFGLTSLQVVGVIAWLGAQPPHSVIDYEEQRTVDPEQARGVLKCDMSDLSLIGCLGYSLLLMVTCTVYAIKARGVPETFNEAKPIGFTMYTTCIIWLAFVPIFFGTAQSAEKIYIQTTTLTVSLSLSASVSLGMLYVPKTYVILFHPEQNVQKRKRSLKKTSTMAAPPQNENAEDAK |
| UniProt | P35349 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | N/A |
| 3D structure model | No available structures or models |
| BioLiP | N/A |
| Therapeutic Target Database | N/A |
| ChEMBL | CHEMBL3842 |
| IUPHAR | 294 |
| DrugBank | N/A |
| Name | CHEMBL262416 |
|---|---|
| Molecular formula | C8H12N2O4 |
| IUPAC name | (2R)-2-amino-4-(5-methyl-3-oxo-1,2-oxazol-4-yl)butanoic acid |
| Molecular weight | 200.194 |
| Hydrogen bond acceptor | 5 |
| Hydrogen bond donor | 3 |
| XlogP | -2.9 |
| Synonyms | 2-Amino-4-(3-hydroxy-5-methyl-isoxazol-4-yl)-butyric acid (R)-2-Amino-4-(3-hydroxy-5-methyl-isoxazol-4-yl)-butyric acid (R)-Homo-AMPA BDBM50060718 |
| Inchi Key | NZDIZJGEDFARSV-ZCFIWIBFSA-N |
| Inchi ID | InChI=1S/C8H12N2O4/c1-4-5(7(11)10-14-4)2-3-6(9)8(12)13/h6H,2-3,9H2,1H3,(H,10,11)(H,12,13)/t6-/m1/s1 |
| PubChem CID | 10774389 |
| ChEMBL | CHEMBL262416 |
| IUPHAR | N/A |
| BindingDB | 50060718 |
| DrugBank | N/A |
Structure | 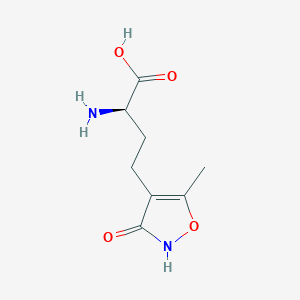 |
| Lipinski's druglikeness | This ligand satisfies Lipinski's rule of five. |
| Parameter | Value | Reference | Database source |
|---|---|---|---|
| EC50 | <1000000.0 nM | PMID9357538 | BindingDB,ChEMBL |
zhanglab![]() zhanggroup.org | +65-6601-1241 | Computing 1, 13 Computing Drive, Singapore 117417
zhanggroup.org | +65-6601-1241 | Computing 1, 13 Computing Drive, Singapore 117417