| Synonyms | HY-17498
Q-200656
InChI=1/C14H22N2O3/c1-10(2)16-8-12(17)9-19-13-5-3-11(4-6-13)7-14(15)18/h3-6,10,12,16-17H,7-9H2,1-2H3,(H2,15,18)
Seles beta
KBio1_000057
Spectrum3_001448
KS-000046IQ
CAS-29122-68-7
LP00121
CS-2927
MLS001066372
dl-Atenolol
NCGC00015007-07
EINECS 262-544-7
NCGC00024566-05
GEO-03413
Normiten
HMS3259K08
Panapres
Atenolol, United States Pharmacopeia (USP) Reference Standard
60966-51-0
BBC/163
AC1Q1QBX
benzeneacetamide, 4-[2'-hydroxy-3;-[(1-methylethyl)amino]propoxy]-
Anselol
Blocotenol
Atcardil
SR-01000000159-8
Cardiopress
Atenol
Tenoblock
(+-)-4-(2-Hydroxy-3-isopropylaminopropoxy)phenylacetamide
Atenol Gador
Tensimin
(inverted question mark)-Atenolol
Atenol Quesada
Vascoten
122A687
Atenolin
2-(p-(2-Hydroxy-3-(isopropylamino)propoxy)phenyl)acetamide
Atenolol [USAN:BAN:INN:JAN]
Ibinolo
SBB056998
Jenatenol
Servitenol
KBio2_006980
L000116
CHEBI:2904
MCULE-9148694071
D00235
MolPort-001-792-717
DSSTox_RID_76663
NCGC00015007-10
Evitocor
NCGC00255122-01
HMS1569L13
NSC757832
HMS3369B14
Plenacor
Atenololum [INN-Latin]
AB00052208-15
BC201805
Aircrit
Betacard
API0001553
Spectrum_001364
BRD-A20239487-001-15-7
Atenblock
Stermin
Atenol AL
Tenoretic (Salt/Mix)
(+/-)-4-[2-Hydroxy-3-[(1-methylethyl)amino]propoxy]benzeneacetamide
Atenol Heumann
Tox21_500121
(RS)-Atenolol
Atenol Trom
Xaten
2-(4-(2-Hydroxy-3-isopropylaminopropoxy)phenyl)acetamid
Atenolol (JP15/USP/INN)
2-[4-[2-hydroxy-3-(isopropylamino)propoxy]phenyl]acetamide
Atenolol, 98%
4-[2 -Hydroxy-3 -(isopropylamino)-propoxy]phenylacetamide
Prenormine
ICI-66082
Scheinpharm Atenol
KB-47450
SPECTRUM1501127
KBioSS_001844
Loten
Corotenol
MFCD00057645
DB00335
NC00548
duratenol
NCGC00024566-03
FT-0622499
NINDS_000057
HMS2092D19
Oraday
HMS500C19
4-[2-Hydroxy-3-(isopropylamino)-propoxy]phenylacetamide
Aterol
AC-8245
Benzeneacetamide, 4-(2-hydroxy-3-((1-methylethyl)amino)propoxy)-
Altol
BG0049
Artrenolol
SR-01000000159-4
CA0145
Ateni
Teno-basan
( inverted question mark)-Atenolol
Atenol ct
Tenormine
(?)-Atenolol
Atenol Nordic
Unibloc
1-p-Carbamoylmethylphenoxy-3-isopropylamino-2-propanol
Atenol-ratiopharm
ZX-AS004380
2-(4-{2-hydroxy-3-[(propan-2-yl)amino]propoxy}phenyl)acetamide
Atenolol Bp
2-{4-[2-hydroxy-3-(isopropylamino)propoxy]phenyl}acetamide
Atenolol, European Pharmacopoeia (EP) Reference Standard
Hypoten
RTR-012731
Internolol
Selobloc
KBio2_001844
Spectrum4_000435
KS-5341
CCG-39010
LS-28557
CTK3J2163
MLS001074163
DSSTox_CID_2628
NCGC00015007-08
EN300-119532
NCGC00024566-06
GTPL548
Noten
HMS3260I04
Pharmakon1600-01501127
Atenolol,(S)
A 7655
BBL009276
Acetamide, 2-(p-(2-hydroxy-3-(isopropylamino)propoxy)phenyl)-
Benzeneacetamide, 4-[2-hydroxy-3-[(1-methylethyl)amino]propoxy]-
Antipressan
Blokium
Atecard
ST024754
Atenol 1A pharma
Tenolol
(+-)-Atenolol
Atenol Genericon
Tensotin
(r,s)-atenolol
Atenol Stada
Vericordin
2-(4-(2-hydroxy-3-(isopropylamino)propoxy)
2-[4-(2-Hydroxy-3-isopropylaminopropoxy)phenyl]acetamide
Atenolol [USAN:INN:BAN:JAN]
29122-68-7
Premorine
ICI 66,082
SBI-0050109.P003
Juvental
SMR000036768
KBio3_002415
Lo-ten
METKIMKYRPQLGS-UHFFFAOYSA-
D01UXC
MRF-0000571
DTXSID2022628
NCGC00015007-11
Farnormin
NCGC00260806-01
HMS1921H09
Opera_ID_1283
HMS3369D20
4-[2'-Hydroxy-3'-(isopropylamino)-propoxy]phenylacetamide
Atenomel
AB00052208_16
BCP12899
AKOS005111050
Betasyn
Apo-Atenolol
SR-01000000159
BRN 2739235
Atendol
STK528649
Atenol Atid
Tenormin
(+/-)-Atenolol
Atenol MSD
TR-012731
(y)-Atenolol
Atenol von ct
Z-0828
2-(4-[2-Hydroxy-3-(isopropylamino)propoxy]phenyl)acetamide
Atenolol (JP17/USP/INN)
2-[4-[2-hydroxy-3-(propan-2-ylamino)propoxy]phenyl]acetamide
Atenolol, >=98% (TLC), powder
Prinorm
IDI1_000057
SCHEMBL4362
KB-62479
Spectrum2_001411
KS-00000XIG
Lotenal
CPD000036768
MLS000069622
DivK1c_000057
NCGC00015007-06
EINECS 249-451-7
NCGC00024566-04
FT-0693045
Normalol
HMS2233E06
Ormidol
HSDB 6526
AX8005715
AC1L1D99
BENZENEACETAMIDE, 4-(2-HYDROXY-3-((1-METHYLETHYL)AMINO)PROPOXY)-, (+-)-
AN-7253
BIM-0050109.0001
Atcard
SR-01000000159-5
Cardaxen
Atenil
Tenobloc
(+)-4-[2-Hydroxy-3-[(1-methylethyl)amino]propoxy]benzeneacetamide
Atenol Fecofar
Tenormine [French]
(A+/-)-Atenolol
Atenol PB
Uniloc
106020-65-9
Atenol-Wolff
2-(4-{[(2S)-2-hydroxy-3-(propan-2-ylamino)propyl]oxy}phenyl)acetamide
Atenolol solution, 1.0 mg/mL in acetonitrile, ampule of 1 mL, certified reference material
2-{4-[2-hydroxy-3-(propan-2-ylamino)propoxy]phenyl}acetamide
Atenolol, Pharmaceutical Secondary Standard; Certified Reference Material
I01-3572
SAM002564193
J10009
Serten
KBio2_004412
Spectrum5_001509
KSC492C6H
CCRIS 4196
LS-9707
Cuxanorm
MLS001304038
DSSTox_GSID_22628
NCGC00015007-09
EU-0100121
NCGC00024566-07
Hipres
NSC-757832
HMS3266K13
phenyl)acetamide
Atenololum
AB00052208-13
BB_SC-01519
Acetamide,2-(p-(2-hydroxy-3-(isopropylamino)propoxy)phenyl)
Betablok
ANW-45215
BRD-A20239487-001-02-5
Atehexal
ST2403914
Atenol acis
Tenoprin
(+/-)-4-(2-Hydroxy-3-[(1-methylethyl)amino]propoxy)benzeneacetamide
Atenol GNR
Tox21_302426
(RS)-4-[2-Hydroxy-3-[(1-methylethyl)amino]propoxy]benzeneacetamide
Atenol Tika
Wesipin
2-(4-(2-hydroxy-3-(isopropylamino)propoxy)phenyl)acetamide
Atenolol (JAN/USP)
2-[4-({2-hydroxy-3-[(1-methylethyl)amino]propyl}oxy)phenyl]acetamide
Atenolol [USAN:USP:INN:BAN:JAN]
4-(2-Hydroxy-3-((1-methylethyl)amino)propoxy)benzeneacetamide
Prenolol
ICI 66082
SC-19545
KB-222053
SPBio_001482
KBioGR_000790
Lopac0_000121
CHEMBL24
METKIMKYRPQLGS-UHFFFAOYSA-N
Myocord
Duraatenolol
NCGC00015007-13
Felo-Bits
Neatenol
HMS2090I19
Oprea1_448775
HMS3369P20
4-[2'-Hydroxy-3'-(isopropylamino)propoxy]phenylacetamide
Atereal
AB0011597
BDBM25753
Alinor
Betatop Ge
ARONIS23884
SR-01000000159-2
BSPBio_002915
Atenet
Tenidon
Atenol Cophar
Tenormin (TN)
(1)-2-(4-(2-Hydroxy-3-(isopropylamino)propoxy)phenyl)acetamide
Atenol NM Pharma
Tredol
Atenol-Mepha
Z1541638518
2-(4-{2-hydroxy-3-[(methylethyl)amino]propoxy}phenyl)acetamide
Atenolol 1.0 mg/ml in Acetonitrile
2-{4-[2-Hydroxy-3-(isopropylamino)propoxy]-phenyl}acetamide
Atenolol, analytical reference material [ Show all ] |
|---|
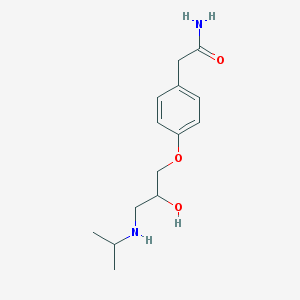
| Inchi ID | InChI=1S/C14H22N2O3/c1-10(2)16-8-12(17)9-19-13-5-3-11(4-6-13)7-14(15)18/h3-6,10,12,16-17H,7-9H2,1-2H3,(H2,15,18) |
|---|


![]() zhanggroup.org | +65-6601-1241 | Computing 1, 13 Computing Drive, Singapore 117417
zhanggroup.org | +65-6601-1241 | Computing 1, 13 Computing Drive, Singapore 117417