

You can:
| Name | Neuropeptide S receptor |
|---|---|
| Species | Mus musculus (Mouse) |
| Gene | Npsr1 |
| Synonym | vasopressin receptor-related receptor 1 PGR14 NPS receptor GPR154 G-protein coupled receptor PGR14 [ Show all ] |
| Disease | N/A for non-human GPCRs |
| Length | 371 |
| Amino acid sequence | MPANLTEGSFHANQTVPMLDSSPVACTEIVTFTEALVAEEWGSFYSSFKTEQLITLWVLFVVTIVGNSVVLFSTCRRKRKSRMTFFVTQLAITDSFTGLINILTDIIWRFTGDFMAPDLVCRVVRYLQVVLLYASTYVLVSLSIDRYHAIVYPMKFLQGEKQAKVLIGIAWSLSFLFSIPTLIIFGKRTLSNGEVQCWALWPDDSYWTPYMTIVAFLVYFIPLAIISVIYGLVIRTIWMKSKTHETVISNCSDGKLCCSYNRGLISKAKIKAIKYSIVIILAFICCWSPYFLFDILDNFNVLPDTKERFYASVIIQNLPALNSAINPLIYCIFSSSICSPCKMQRSQDSRMTYRERSERHEMQILSKPEFI |
| UniProt | Q8BZP8 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | N/A |
| 3D structure model | No available structures or models |
| BioLiP | N/A |
| Therapeutic Target Database | N/A |
| ChEMBL | CHEMBL5497 |
| IUPHAR | 302 |
| DrugBank | N/A |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 1756 | 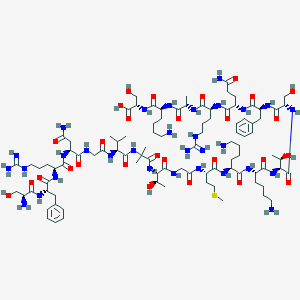 CHEMBL266339 CHEMBL266339 | C95H159N31O28S | 2215.57 | 35 / 37 | -13.1 | No |
| 11345 | 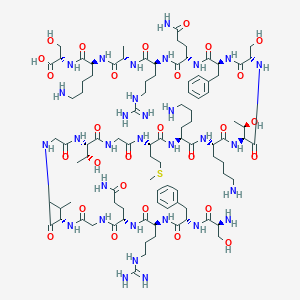 CHEMBL451444 CHEMBL451444 | C94H157N31O28S | 2201.54 | 35 / 37 | -13.3 | No |
| 11971 | 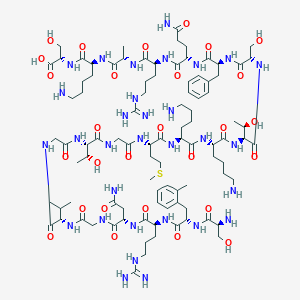 CHEMBL398483 CHEMBL398483 | C94H157N31O28S | 2201.54 | 35 / 37 | -13.3 | No |
| 16393 | 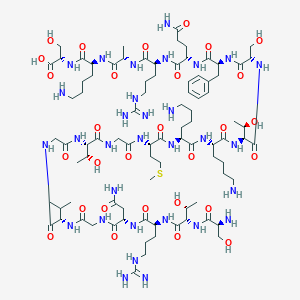 CHEMBL409556 CHEMBL409556 | C88H153N31O29S | 2141.44 | 36 / 38 | -15.9 | No |
| 16817 | 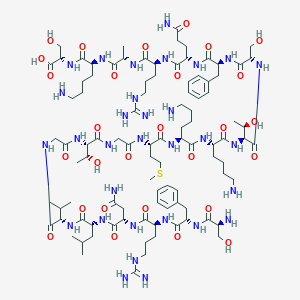 CHEMBL525809 CHEMBL525809 | C97H163N31O28S | 2243.62 | 35 / 37 | -11.9 | No |
| 16818 | 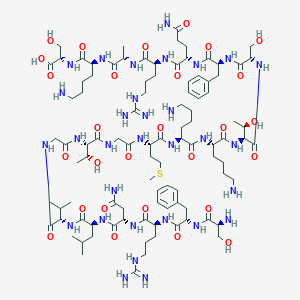 CHEMBL526323 CHEMBL526323 | C97H163N31O28S | 2243.62 | 35 / 37 | -11.9 | No |
| 20571 | 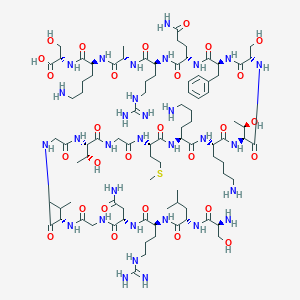 CHEMBL268436 CHEMBL268436 | C90H157N31O28S | 2153.5 | 35 / 37 | -13.9 | No |
| 21508 | 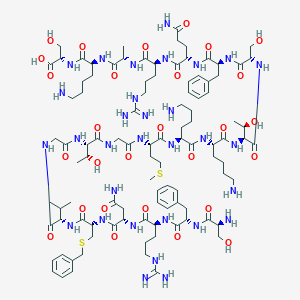 CHEMBL524649 CHEMBL524649 | C101H163N31O28S2 | 2323.72 | 36 / 37 | -11.5 | No |
| 22420 | 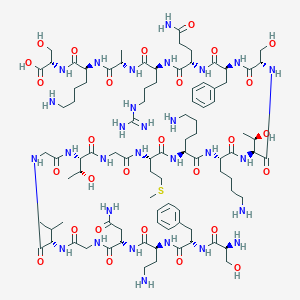 CHEMBL510673 CHEMBL510673 | C91H151N29O28S | 2131.44 | 35 / 35 | -13.8 | No |
| 24387 | 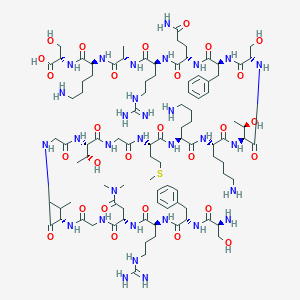 CHEMBL504006 CHEMBL504006 | C95H159N31O28S | 2215.57 | 35 / 36 | -13.1 | No |
| 24747 | 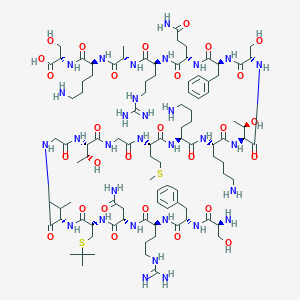 CHEMBL507041 CHEMBL507041 | C98H165N31O28S2 | 2289.71 | 36 / 37 | -12.0 | No |
| 34785 | 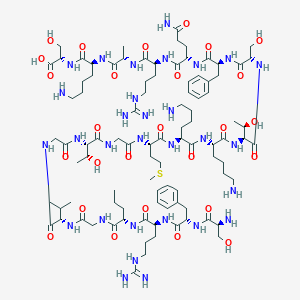 CHEMBL525197 CHEMBL525197 | C94H158N30O27S | 2172.54 | 34 / 36 | -11.2 | No |
| 35236 | 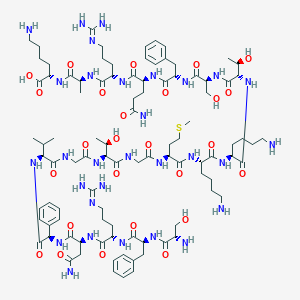 BDBM50413779 BDBM50413779 | C96H154N30O26S | 2176.53 | 33 / 33 | -11.4 | No |
| 39674 | 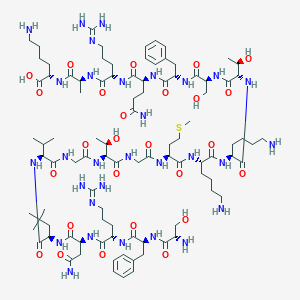 BDBM50413778 BDBM50413778 | C95H160N30O26S | 2170.57 | 33 / 33 | -11.0 | No |
| 43582 | 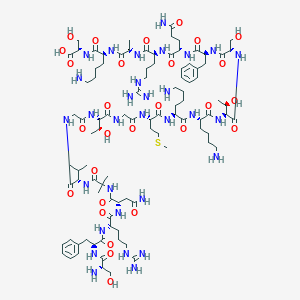 CHEMBL267479 CHEMBL267479 | C95H159N31O28S | 2215.57 | 35 / 37 | -13.1 | No |
| 63226 | 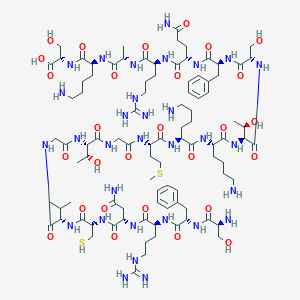 CID 25193412 CID 25193412 | C94H157N31O28S2 | 2233.6 | 36 / 38 | -13.3 | No |
| 63227 | 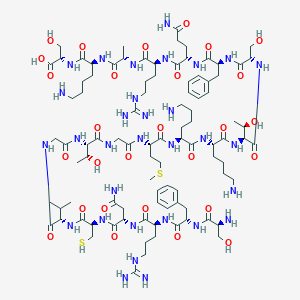 CHEMBL526355 CHEMBL526355 | C94H157N31O28S2 | 2233.6 | 36 / 38 | -13.3 | No |
| 66578 | 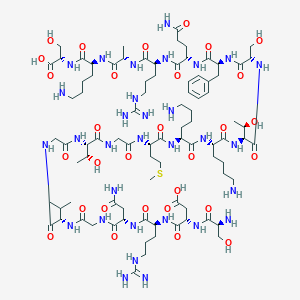 CHEMBL398849 CHEMBL398849 | C88H151N31O30S | 2155.42 | 37 / 38 | -18.4 | No |
| 84308 | 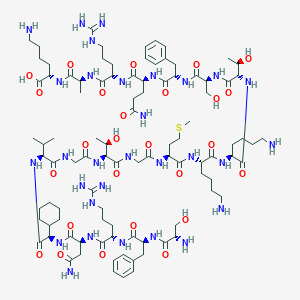 CHEMBL525618 CHEMBL525618 | C96H160N30O26S | 2182.58 | 33 / 35 | -9.9 | No |
| 85119 | 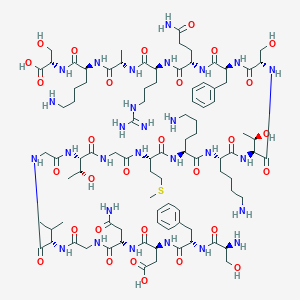 CHEMBL504008 CHEMBL504008 | C91H148N28O30S | 2146.41 | 36 / 35 | -16.0 | No |
| 88140 | 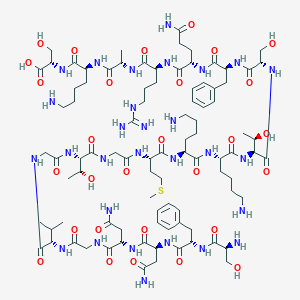 CHEMBL499993 CHEMBL499993 | C91H149N29O29S | 2145.43 | 35 / 35 | -14.3 | No |
| 92015 | 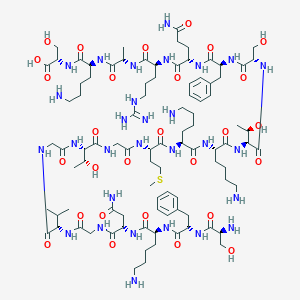 CHEMBL526551 CHEMBL526551 | C93H155N29O28S | 2159.5 | 35 / 35 | -13.1 | No |
| 102316 | 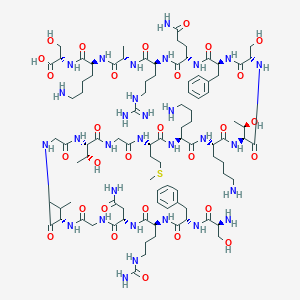 CHEMBL447434 CHEMBL447434 | C93H154N30O29S | 2188.5 | 35 / 36 | -13.7 | No |
| 110208 | 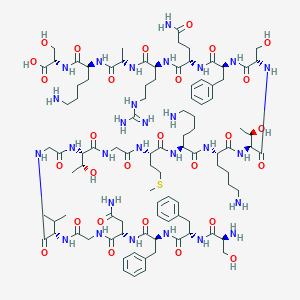 CHEMBL505840 CHEMBL505840 | C96H152N28O28S | 2178.5 | 34 / 34 | -11.2 | No |
| 113800 | 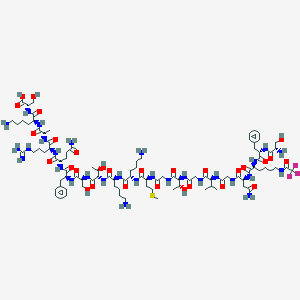 CHEMBL525031 CHEMBL525031 | C95H154F3N29O29S | 2255.51 | 38 / 35 | -11.7 | No |
| 129472 | 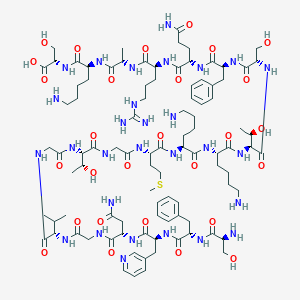 CHEMBL526137 CHEMBL526137 | C95H151N29O28S | 2179.49 | 35 / 34 | -12.3 | No |
| 133974 | 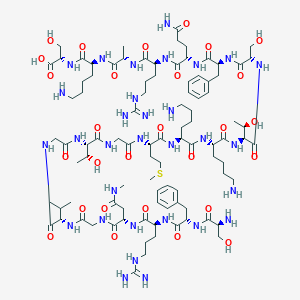 CHEMBL510921 CHEMBL510921 | C94H157N31O28S | 2201.54 | 35 / 37 | -13.2 | No |
| 137114 | 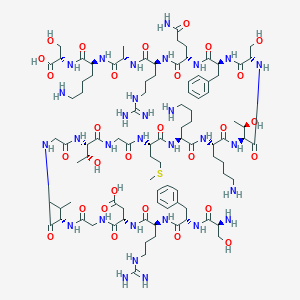 CHEMBL452889 CHEMBL452889 | C93H154N30O29S | 2188.5 | 36 / 37 | -15.3 | No |
| 145918 | 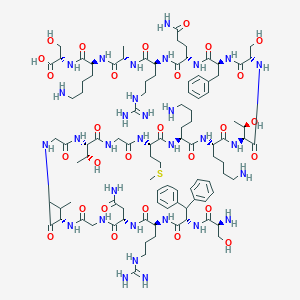 CHEMBL436829 CHEMBL436829 | C99H159N31O28S | 2263.61 | 35 / 37 | -12.2 | No |
| 158956 | 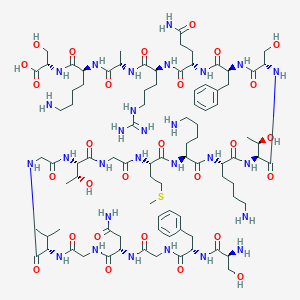 CHEMBL499957 CHEMBL499957 | C89H146N28O28S | 2088.38 | 34 / 34 | -13.2 | No |
| 163271 | 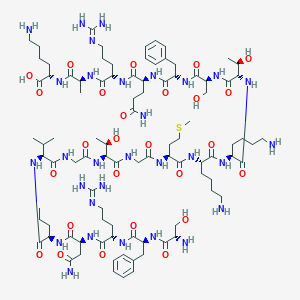 BDBM50413780 BDBM50413780 | C93H156N30O26S | 2142.51 | 33 / 33 | -11.8 | No |
| 166077 | 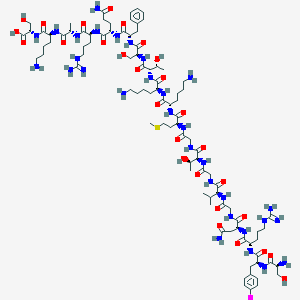 CHEMBL267034 CHEMBL267034 | C93H154IN31O28S | 2313.41 | 35 / 37 | -13.0 | No |
| 169675 | 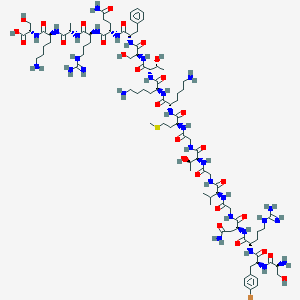 CHEMBL438162 CHEMBL438162 | C93H154BrN31O28S | 2266.41 | 35 / 37 | -13.0 | No |
| 175564 | 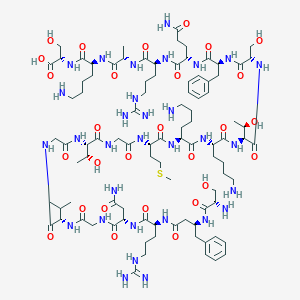 CHEMBL439237 CHEMBL439237 | C94H157N31O28S | 2201.54 | 35 / 37 | -13.7 | No |
| 177714 | 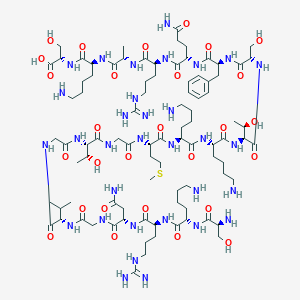 CHEMBL429312 CHEMBL429312 | C90H158N32O28S | 2168.51 | 36 / 38 | -15.5 | No |
| 177894 | 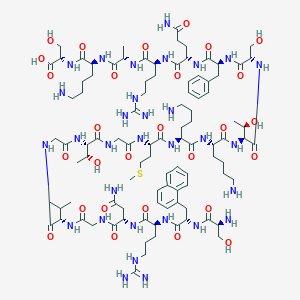 CHEMBL398524 CHEMBL398524 | C97H157N31O28S | 2237.57 | 35 / 37 | -12.4 | No |
| 186559 | 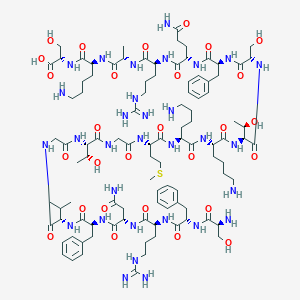 CHEMBL525398 CHEMBL525398 | C100H161N31O28S | 2277.64 | 35 / 37 | -11.7 | No |
| 186560 | 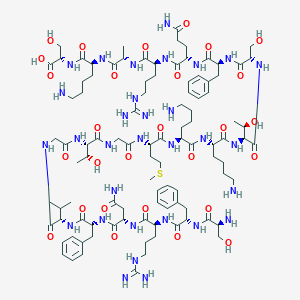 CHEMBL508167 CHEMBL508167 | C100H161N31O28S | 2277.64 | 35 / 37 | -11.7 | No |
| 187981 | 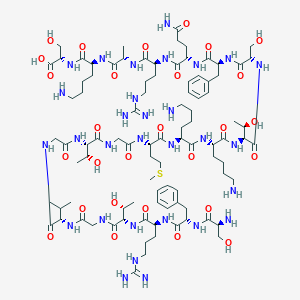 CHEMBL453962 CHEMBL453962 | C93H156N30O28S | 2174.51 | 35 / 37 | -12.7 | No |
| 189283 | 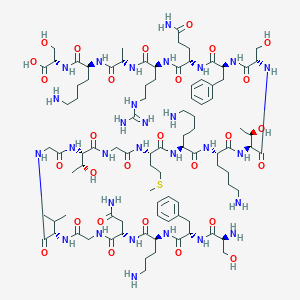 CHEMBL510338 CHEMBL510338 | C92H153N29O28S | 2145.47 | 35 / 35 | -13.4 | No |
| 190368 | 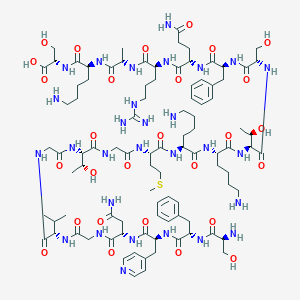 CHEMBL500037 CHEMBL500037 | C95H151N29O28S | 2179.49 | 35 / 34 | -12.3 | No |
| 191399 | 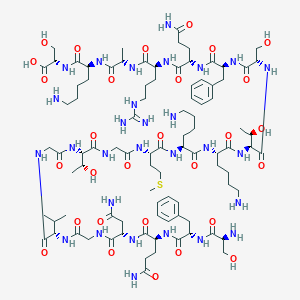 CHEMBL506383 CHEMBL506383 | C92H151N29O29S | 2159.45 | 35 / 35 | -14.0 | No |
| 191470 | 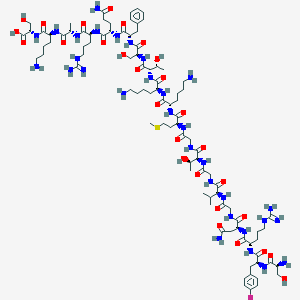 CHEMBL430024 CHEMBL430024 | C93H154FN31O28S | 2205.5 | 36 / 37 | -13.6 | No |
| 203871 | 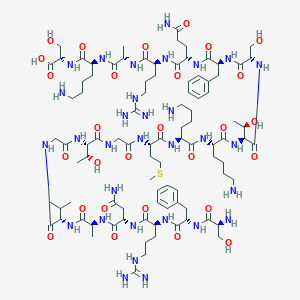 CHEMBL438115 CHEMBL438115 | C94H157N31O28S | 2201.54 | 35 / 37 | -13.2 | No |
| 203872 | 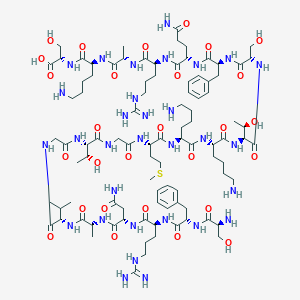 CHEMBL263801 CHEMBL263801 | C94H157N31O28S | 2201.54 | 35 / 37 | -13.2 | No |
| 204289 | 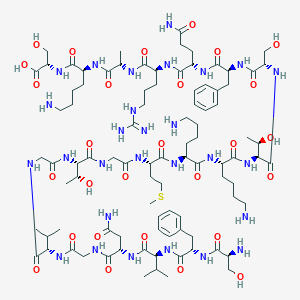 CHEMBL525271 CHEMBL525271 | C92H152N28O28S | 2130.46 | 34 / 34 | -11.9 | No |
| 211333 | 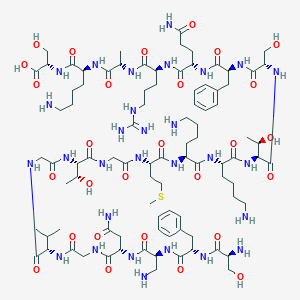 CHEMBL505996 CHEMBL505996 | C90H149N29O28S | 2117.42 | 35 / 35 | -14.1 | No |
| 213465 | 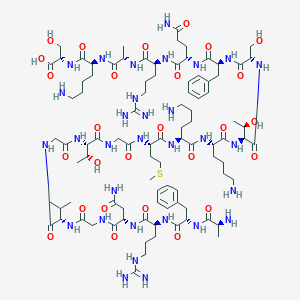 CHEMBL229994 CHEMBL229994 | C93H155N31O27S | 2171.51 | 34 / 36 | -12.6 | No |
| 215798 | 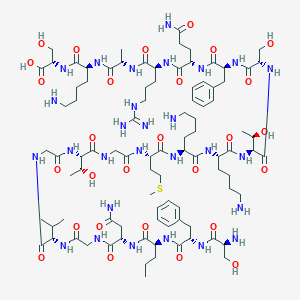 CHEMBL525447 CHEMBL525447 | C92H152N28O28S | 2130.46 | 34 / 34 | -11.9 | No |
| 220044 | 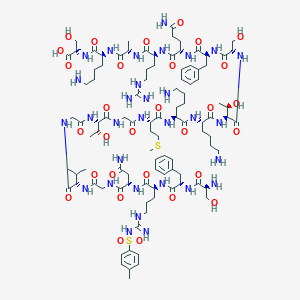 CHEMBL525806 CHEMBL525806 | C100H161N31O30S2 | 2341.7 | 37 / 37 | -11.8 | No |
| 225901 | 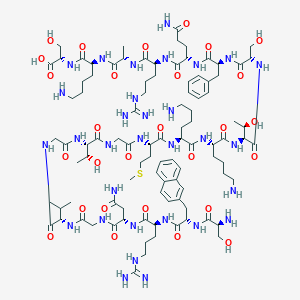 [2Nal2]hNPS [2Nal2]hNPS | C97H157N31O28S | 2237.57 | 35 / 37 | -12.4 | No |
| 226847 | 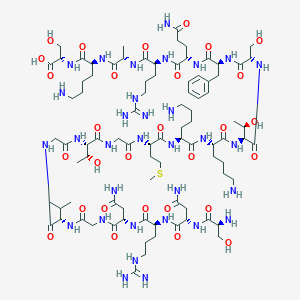 CHEMBL251926 CHEMBL251926 | C88H152N32O29S | 2154.44 | 36 / 38 | -16.8 | No |
| 231966 | 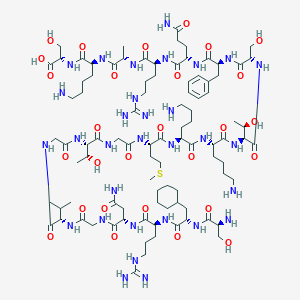 CHEMBL409981 CHEMBL409981 | C93H161N31O28S | 2193.56 | 35 / 37 | -12.5 | No |
| 233450 | 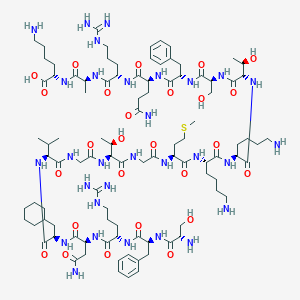 CHEMBL526895 CHEMBL526895 | C97H162N30O26S | 2196.61 | 33 / 35 | -9.2 | No |
| 243026 | 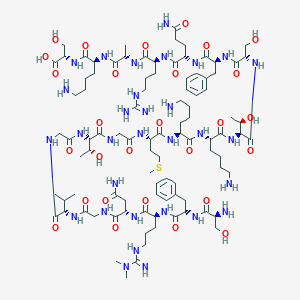 CHEMBL524654 CHEMBL524654 | C95H159N31O28S | 2215.57 | 35 / 36 | -13.1 | No |
| 248060 | 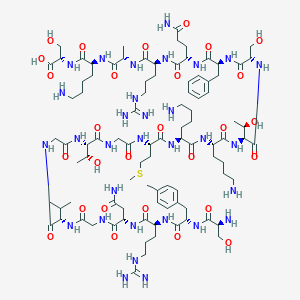 CHEMBL437201 CHEMBL437201 | C94H157N31O28S | 2201.54 | 35 / 37 | -13.3 | No |
| 253905 | 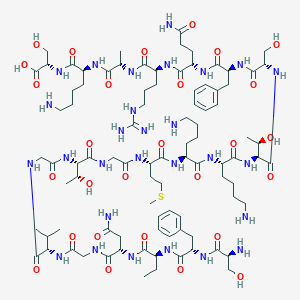 CHEMBL505174 CHEMBL505174 | C91H150N28O28S | 2116.43 | 34 / 34 | -12.3 | No |
| 256645 | 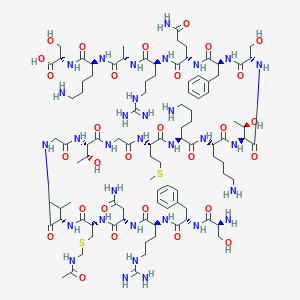 CHEMBL508019 CHEMBL508019 | C97H162N32O29S2 | 2304.68 | 37 / 38 | -13.6 | No |
| 258786 | 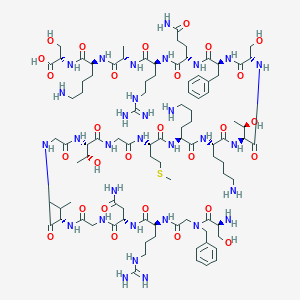 CHEMBL409555 CHEMBL409555 | C93H155N31O28S | 2187.51 | 35 / 36 | -14.0 | No |
| 261145 | 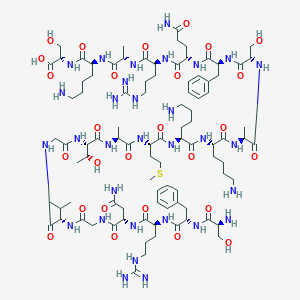 CHEMBL437838 CHEMBL437838 | C93H155N31O27S | 2171.51 | 34 / 36 | -12.6 | No |
| 266837 | 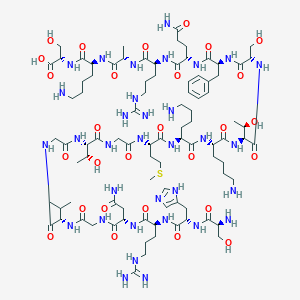 CHEMBL401641 CHEMBL401641 | C90H153N33O28S | 2177.48 | 36 / 38 | -15.5 | No |
| 267503 | 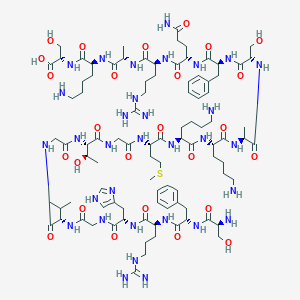 CHEMBL2372037 CHEMBL2372037 | C94H154N32O26S | 2180.52 | 34 / 36 | -11.7 | No |
| 276238 | 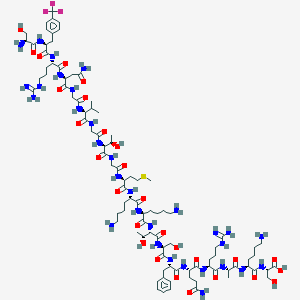 CHEMBL437389 CHEMBL437389 | C94H154F3N31O28S | 2255.51 | 38 / 37 | -12.8 | No |
| 279475 | 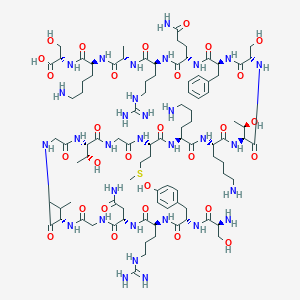 CHEMBL437199 CHEMBL437199 | C93H155N31O29S | 2203.51 | 36 / 38 | -14.0 | No |
| 280993 | 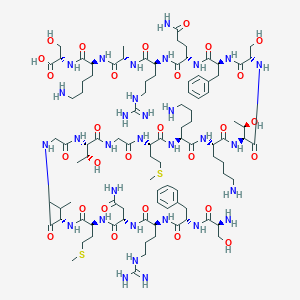 CHEMBL507881 CHEMBL507881 | C96H161N31O28S2 | 2261.65 | 36 / 37 | -12.6 | No |
| 280994 | 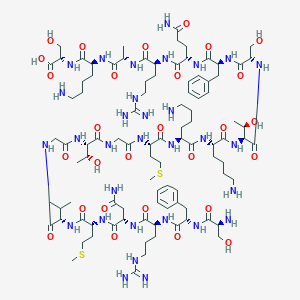 CHEMBL524496 CHEMBL524496 | C96H161N31O28S2 | 2261.65 | 36 / 37 | -12.6 | No |
| 282011 | 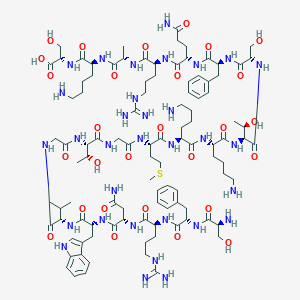 CHEMBL525797 CHEMBL525797 | C102H162N32O28S | 2316.67 | 35 / 38 | -11.5 | No |
| 282012 | 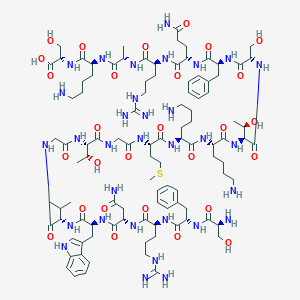 CHEMBL498862 CHEMBL498862 | C102H162N32O28S | 2316.67 | 35 / 38 | -11.5 | No |
| 289879 | 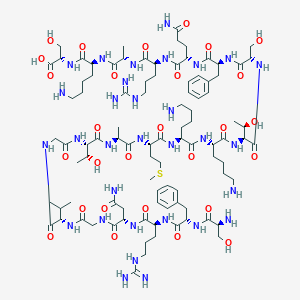 CHEMBL438866 CHEMBL438866 | C94H157N31O28S | 2201.54 | 35 / 37 | -13.2 | No |
| 289880 | 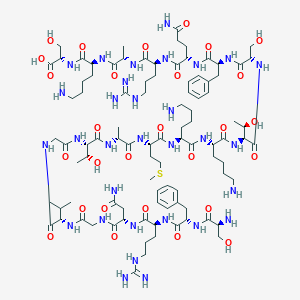 CHEMBL230108 CHEMBL230108 | C94H157N31O28S | 2201.54 | 35 / 37 | -13.2 | No |
| 296466 | 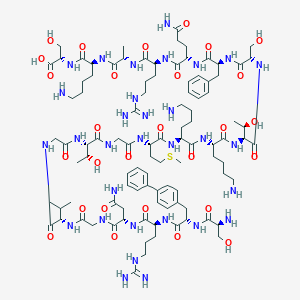 CHEMBL267266 CHEMBL267266 | C99H159N31O28S | 2263.61 | 35 / 37 | -12.0 | No |
| 299974 | 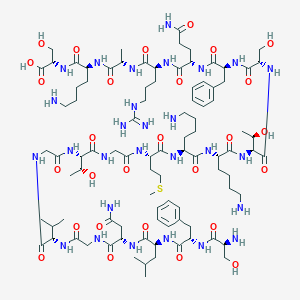 CHEMBL524536 CHEMBL524536 | C93H154N28O28S | 2144.48 | 34 / 34 | -11.5 | No |
| 303030 | 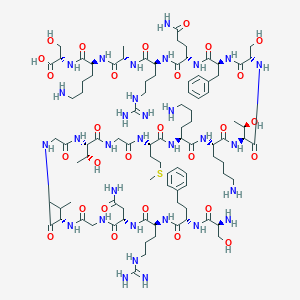 CHEMBL413849 CHEMBL413849 | C94H157N31O28S | 2201.54 | 35 / 37 | -13.3 | No |
| 303412 |  CHEMBL441576 CHEMBL441576 | C93H154ClN31O28S | 2221.95 | 35 / 37 | -13.0 | No |
| 313167 |  CHEMBL1765703 CHEMBL1765703 | C26H24FN3O3 | 445.494 | 4 / 1 | 3.7 | Yes |
| 313169 | 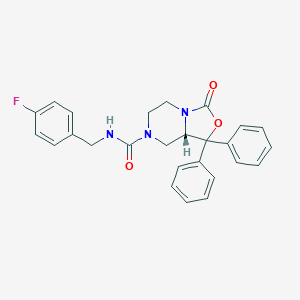 CHEMBL1765704 CHEMBL1765704 | C26H24FN3O3 | 445.494 | 4 / 1 | 3.7 | Yes |
| 313171 | 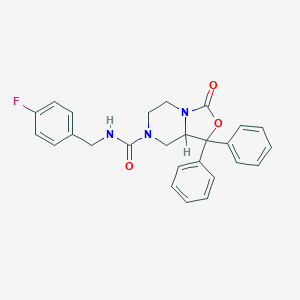 SHA-68 SHA-68 | C26H24FN3O3 | 445.494 | 4 / 1 | 3.7 | Yes |
| 326580 | 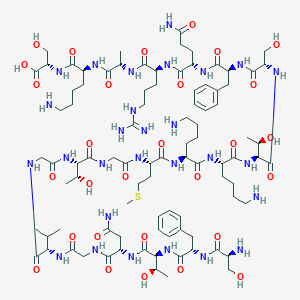 CHEMBL503354 CHEMBL503354 | C91H150N28O29S | 2132.43 | 35 / 35 | -13.4 | No |
| 329957 | 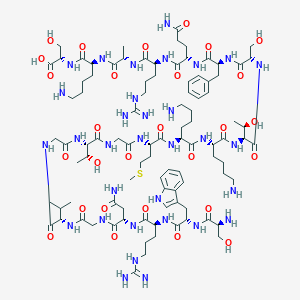 CHEMBL400369 CHEMBL400369 | C95H156N32O28S | 2226.55 | 35 / 38 | -13.5 | No |
| 331648 | 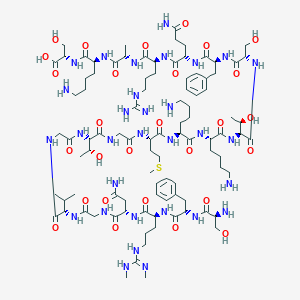 CHEMBL524688 CHEMBL524688 | C95H159N31O28S | 2215.57 | 35 / 36 | -13.2 | No |
| 335744 | 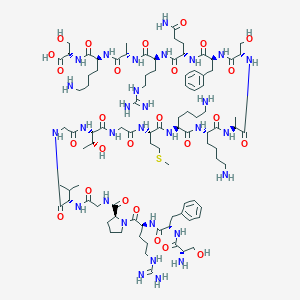 CHEMBL396531 CHEMBL396531 | C93H154N30O26S | 2140.5 | 33 / 34 | -11.2 | No |
| 335745 | 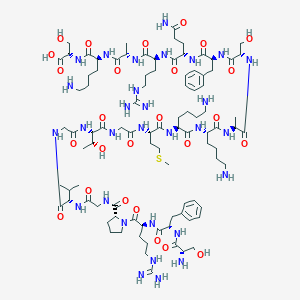 CHEMBL440059 CHEMBL440059 | C93H154N30O26S | 2140.5 | 33 / 34 | -11.2 | No |
| 336239 | 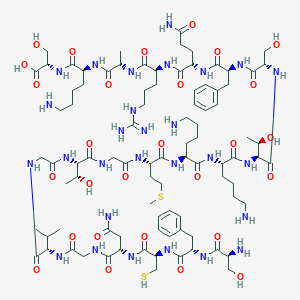 CHEMBL503441 CHEMBL503441 | C90H148N28O28S2 | 2134.46 | 35 / 35 | -12.9 | No |
| 336499 | 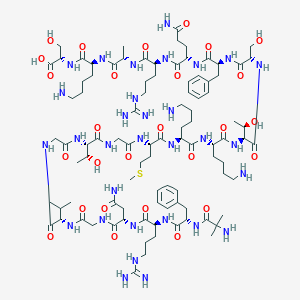 CHEMBL425840 CHEMBL425840 | C94H157N31O27S | 2185.54 | 34 / 36 | -12.4 | No |
| 338385 | 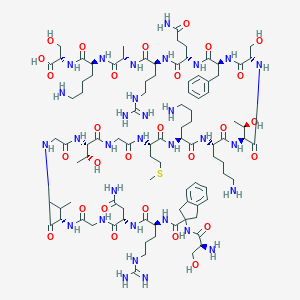 CHEMBL411571 CHEMBL411571 | C94H155N31O28S | 2199.52 | 35 / 37 | -13.2 | No |
| 338610 | 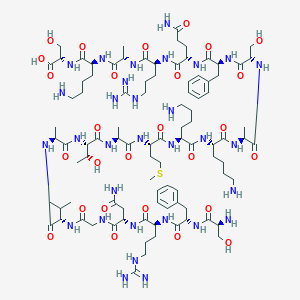 CHEMBL406582 CHEMBL406582 | C94H157N31O27S | 2185.54 | 34 / 36 | -12.2 | No |
| 340541 | 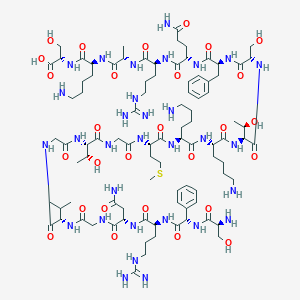 CHEMBL437281 CHEMBL437281 | C92H153N31O28S | 2173.49 | 35 / 37 | -13.9 | No |
| 344262 | 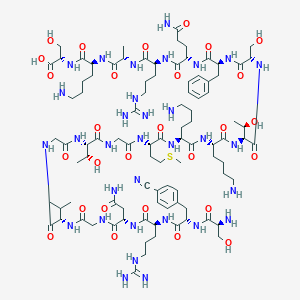 CHEMBL252537 CHEMBL252537 | C94H154N32O28S | 2212.52 | 36 / 37 | -13.9 | No |
| 344772 |  CHEMBL527093 CHEMBL527093 | C93H156N30O26S2 | 2174.57 | 34 / 36 | -11.4 | No |
| 358804 | 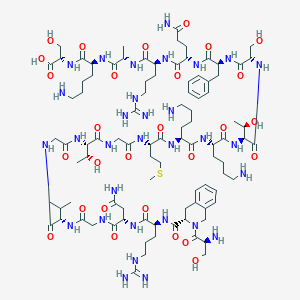 CHEMBL427638 CHEMBL427638 | C94H155N31O28S | 2199.52 | 35 / 36 | -13.9 | No |
| 360492 | 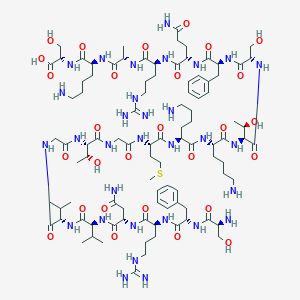 CHEMBL505657 CHEMBL505657 | C96H161N31O28S | 2229.59 | 35 / 37 | -12.3 | No |
| 360493 | 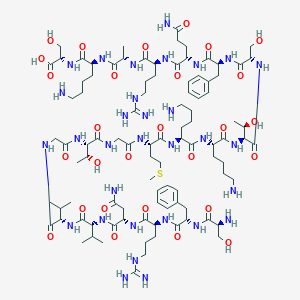 CHEMBL525750 CHEMBL525750 | C96H161N31O28S | 2229.59 | 35 / 37 | -12.3 | No |
| 362510 | 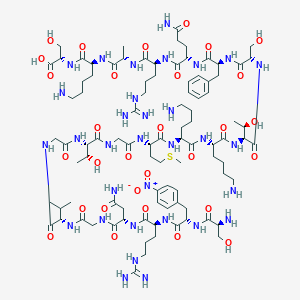 CHEMBL411392 CHEMBL411392 | C93H154N32O30S | 2232.51 | 37 / 37 | -13.8 | No |
| 366952 | 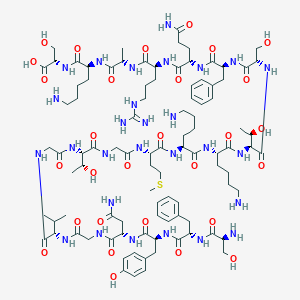 CHEMBL449744 CHEMBL449744 | C96H152N28O29S | 2194.5 | 35 / 35 | -11.6 | No |
| 371845 | 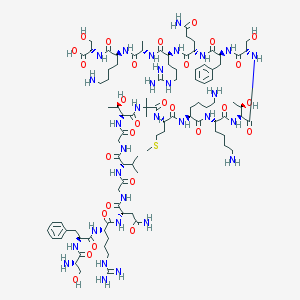 CHEMBL390860 CHEMBL390860 | C95H159N31O28S | 2215.57 | 35 / 37 | -13.1 | No |
| 372169 |  CID 44186330 CID 44186330 | C97H163N31O28S | 2243.62 | 35 / 37 | -11.9 | No |
| 378201 |  CHEMBL399048 CHEMBL399048 | C89H154N32O29S | 2168.47 | 36 / 38 | -16.4 | No |
| 379403 |  CHEMBL484472 CHEMBL484472 | C94H156N30O28S | 2186.52 | 35 / 36 | -12.8 | No |
| 387684 |  BDBM50418333 BDBM50418333 | C93H156N34O27 | 2182.48 | 34 / 36 | -15.6 | No |
| 392257 |  CHEMBL428331 CHEMBL428331 | C92H153N31O27S | 2157.49 | 34 / 36 | -13.0 | No |
zhanglab![]() zhanggroup.org | +65-6601-1241 | Computing 1, 13 Computing Drive, Singapore 117417
zhanggroup.org | +65-6601-1241 | Computing 1, 13 Computing Drive, Singapore 117417