

You can:
| Name | Trace amine-associated receptor 1 |
|---|---|
| Species | Homo sapiens (Human) |
| Gene | TAAR1 |
| Synonym | trace amine receptor 1 TAR1 TaR-1 TA1 receptor TRAR1 |
| Disease | N/A |
| Length | 339 |
| Amino acid sequence | MMPFCHNIINISCVKNNWSNDVRASLYSLMVLIILTTLVGNLIVIVSISHFKQLHTPTNWLIHSMATVDFLLGCLVMPYSMVRSAEHCWYFGEVFCKIHTSTDIMLSSASIFHLSFISIDRYYAVCDPLRYKAKMNILVICVMIFISWSVPAVFAFGMIFLELNFKGAEEIYYKHVHCRGGCSVFFSKISGVLTFMTSFYIPGSIMLCVYYRIYLIAKEQARLISDANQKLQIGLEMKNGISQSKERKAVKTLGIVMGVFLICWCPFFICTVMDPFLHYIIPPTLNDVLIWFGYLNSTFNPMVYAFFYPWFRKALKMMLFGKIFQKDSSRCKLFLELSS |
| UniProt | Q96RJ0 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | Q96RJ0 |
| 3D structure model | This predicted structure model is from GPCR-EXP Q96RJ0. |
| BioLiP | N/A |
| Therapeutic Target Database | N/A |
| ChEMBL | CHEMBL5857 |
| IUPHAR | 364 |
| DrugBank | BE0001044 |
| Name | Skf-81297 |
|---|---|
| Molecular formula | C16H16ClNO2 |
| IUPAC name | 9-chloro-5-phenyl-2,3,4,5-tetrahydro-1H-3-benzazepine-7,8-diol |
| Molecular weight | 289.759 |
| Hydrogen bond acceptor | 3 |
| Hydrogen bond donor | 3 |
| XlogP | 2.7 |
| Synonyms | 6-Chloro-1-phenyl-2,3,4,5-tetrahydro-1H-benzo[d]azepine-7,8-diol CHEBI:91517 GTPL938 NSC_1218 SKF 81297 hydrochloride [ Show all ] |
| Inchi Key | GHWJEDJMOVUXEC-UHFFFAOYSA-N |
| Inchi ID | InChI=1S/C16H16ClNO2/c17-15-11-6-7-18-9-13(10-4-2-1-3-5-10)12(11)8-14(19)16(15)20/h1-5,8,13,18-20H,6-7,9H2 |
| PubChem CID | 1218 |
| ChEMBL | CHEMBL353335 |
| IUPHAR | 938 |
| BindingDB | 86282 |
| DrugBank | N/A |
Structure | 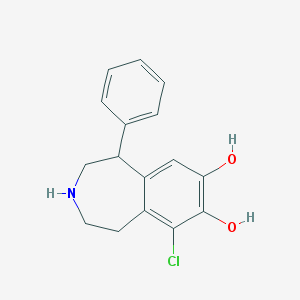 |
| Lipinski's druglikeness | This ligand satisfies Lipinski's rule of five. |
| Parameter | Value | Reference | Database source |
|---|---|---|---|
| EC50 | 6535.0 nM | N/A | BindingDB |
| IC50 | <29913.0 nM | N/A | BindingDB |
zhanglab![]() zhanggroup.org | +65-6601-1241 | Computing 1, 13 Computing Drive, Singapore 117417
zhanggroup.org | +65-6601-1241 | Computing 1, 13 Computing Drive, Singapore 117417