

You can:
| Name | Muscarinic acetylcholine receptor M2 |
|---|---|
| Species | Homo sapiens (Human) |
| Gene | CHRM2 |
| Synonym | cholinergic receptor AChR M2 M2 muscarinic acetylcholine receptor M2 receptor Chrm-2 [ Show all ] |
| Disease | Urinary incontinence Heart failure Nausea; Addiction Parkinson's disease Peptic ulcer [ Show all ] |
| Length | 466 |
| Amino acid sequence | MNNSTNSSNNSLALTSPYKTFEVVFIVLVAGSLSLVTIIGNILVMVSIKVNRHLQTVNNYFLFSLACADLIIGVFSMNLYTLYTVIGYWPLGPVVCDLWLALDYVVSNASVMNLLIISFDRYFCVTKPLTYPVKRTTKMAGMMIAAAWVLSFILWAPAILFWQFIVGVRTVEDGECYIQFFSNAAVTFGTAIAAFYLPVIIMTVLYWHISRASKSRIKKDKKEPVANQDPVSPSLVQGRIVKPNNNNMPSSDDGLEHNKIQNGKAPRDPVTENCVQGEEKESSNDSTSVSAVASNMRDDEITQDENTVSTSLGHSKDENSKQTCIRIGTKTPKSDSCTPTNTTVEVVGSSGQNGDEKQNIVARKIVKMTKQPAKKKPPPSREKKVTRTILAILLAFIITWAPYNVMVLINTFCAPCIPNTVWTIGYWLCYINSTINPACYALCNATFKKTFKHLLMCHYKNIGATR |
| UniProt | P08172 |
| Protein Data Bank | 5zkc, 4mqs, 4mqt, 5yc8, 5zk3, 5zkb, 5zk8 |
| GPCR-HGmod model | P08172 |
| 3D structure model | This structure is from PDB ID 5zkc. |
| BioLiP | BL0433341, BL0433216, BL0263147, BL0263146, BL0263145, BL0433339, BL0433340, BL0433338 |
| Therapeutic Target Database | T46185 |
| ChEMBL | CHEMBL211 |
| IUPHAR | 14 |
| DrugBank | BE0000560 |
| Name | BDBM86295 |
|---|---|
| Molecular formula | C12H16N3O2+ |
| IUPAC name | 2-[4-(2,3-dimethylimidazol-3-ium-1-yl)but-2-ynyl]-1,2-oxazolidin-3-one |
| Molecular weight | 234.279 |
| Hydrogen bond acceptor | 2 |
| Hydrogen bond donor | 0 |
| XlogP | 0.0 |
| Synonyms | PDSP2_001319 2-[4-(2,3-Dimethyl-1H-imidazolium-1-yl)-2-butynyl]isoxazolidin-3-one iodide PDSP1_001335 |
| Inchi Key | AILVCKQCTVDJPU-UHFFFAOYSA-N |
| Inchi ID | InChI=1S/C12H16N3O2/c1-11-13(2)8-9-14(11)6-3-4-7-15-12(16)5-10-17-15/h8-9H,5-7,10H2,1-2H3/q+1 |
| PubChem CID | 23844146 |
| ChEMBL | N/A |
| IUPHAR | N/A |
| BindingDB | 86295 |
| DrugBank | N/A |
Structure | 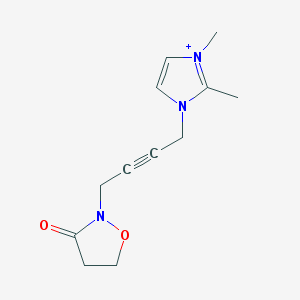 |
| Lipinski's druglikeness | This ligand satisfies Lipinski's rule of five. |
| Parameter | Value | Reference | Database source |
|---|---|---|---|
| Ki | 275.42 nM | PMID13679167 | BindingDB |
zhanglab![]() zhanggroup.org | +65-6601-1241 | Computing 1, 13 Computing Drive, Singapore 117417
zhanggroup.org | +65-6601-1241 | Computing 1, 13 Computing Drive, Singapore 117417