

You can:
| Name | D(4) dopamine receptor |
|---|---|
| Species | Homo sapiens (Human) |
| Gene | DRD4 |
| Synonym | D4 receptor D4R Dopamine D4 receptor dopamine receptor 4 d(2C) dopamine receptor |
| Disease | Erectile dysfunction Psychotic disorders Schizophrenia |
| Length | 467 |
| Amino acid sequence | MGNRSTADADGLLAGRGPAAGASAGASAGLAGQGAAALVGGVLLIGAVLAGNSLVCVSVATERALQTPTNSFIVSLAAADLLLALLVLPLFVYSEVQGGAWLLSPRLCDALMAMDVMLCTASIFNLCAISVDRFVAVAVPLRYNRQGGSRRQLLLIGATWLLSAAVAAPVLCGLNDVRGRDPAVCRLEDRDYVVYSSVCSFFLPCPLMLLLYWATFRGLQRWEVARRAKLHGRAPRRPSGPGPPSPTPPAPRLPQDPCGPDCAPPAPGLPRGPCGPDCAPAAPGLPPDPCGPDCAPPAPGLPQDPCGPDCAPPAPGLPRGPCGPDCAPPAPGLPQDPCGPDCAPPAPGLPPDPCGSNCAPPDAVRAAALPPQTPPQTRRRRRAKITGRERKAMRVLPVVVGAFLLCWTPFFVVHITQALCPACSVPPRLVSAVTWLGYVNSALNPVIYTVFNAEFRNVFRKALRACC |
| UniProt | P21917 |
| Protein Data Bank | 5wiv, 5wiu |
| GPCR-HGmod model | P21917 |
| 3D structure model | This structure is from PDB ID 5wiv. |
| BioLiP | BL0394824, BL0394826, BL0394825 |
| Therapeutic Target Database | T24983 |
| ChEMBL | CHEMBL219 |
| IUPHAR | 217 |
| DrugBank | BE0009376, BE0000389 |
| Name | CHEMBL3983937 |
|---|---|
| Molecular formula | C22H27N5OS |
| IUPAC name | 4-methyl-5-[4-methyl-5-[3-[(2S,3R)-2-phenyl-5-azaspiro[2.4]heptan-5-yl]propylsulfanyl]-1,2,4-triazol-3-yl]-1,3-oxazole |
| Molecular weight | 409.552 |
| Hydrogen bond acceptor | 6 |
| Hydrogen bond donor | 0 |
| XlogP | 3.3 |
| Synonyms | SCHEMBL17712767 BDBM50192034 |
| Inchi Key | AAPXNHMQKBDDJN-PGRDOPGGSA-N |
| Inchi ID | InChI=1S/C22H27N5OS/c1-16-19(28-15-23-16)20-24-25-21(26(20)2)29-12-6-10-27-11-9-22(14-27)13-18(22)17-7-4-3-5-8-17/h3-5,7-8,15,18H,6,9-14H2,1-2H3/t18-,22+/m0/s1 |
| PubChem CID | 121304458 |
| ChEMBL | CHEMBL3983937 |
| IUPHAR | N/A |
| BindingDB | 50192034 |
| DrugBank | N/A |
Structure | 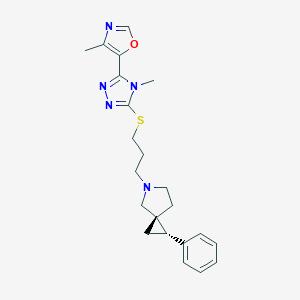 |
| Lipinski's druglikeness | This ligand satisfies Lipinski's rule of five. |
| Parameter | Value | Reference | Database source |
|---|---|---|---|
| Ki | <100000.0 nM | PMID27564135 | ChEMBL |
| Ki | >100000.0 nM | PMID27564135 | BindingDB |
zhanglab![]() zhanggroup.org | +65-6601-1241 | Computing 1, 13 Computing Drive, Singapore 117417
zhanggroup.org | +65-6601-1241 | Computing 1, 13 Computing Drive, Singapore 117417