

You can:
| Name | Trace amine-associated receptor 1 |
|---|---|
| Species | Mus musculus (Mouse) |
| Gene | Taar1 |
| Synonym | TA1 receptor TaR-1 TAR1 trace amine receptor 1 TRAR1 |
| Disease | N/A for non-human GPCRs |
| Length | 332 |
| Amino acid sequence | MHLCHAITNISHRNSDWSREVQASLYSLMSLIILATLVGNLIVIISISHFKQLHTPTNWLLHSMAIVDFLLGCLIMPCSMVRTVERCWYFGEILCKVHTSTDIMLSSASIFHLAFISIDRYCAVCDPLRYKAKINISTILVMILVSWSLPAVYAFGMIFLELNLKGVEELYRSQVSDLGGCSPFFSKVSGVLAFMTSFYIPGSVMLFVYYRIYFIAKGQARSINRTNVQVGLEGKSQAPQSKETKAAKTLGIMVGVFLVCWCPFFLCTVLDPFLGYVIPPSLNDALYWFGYLNSALNPMVYAFFYPWFRRALKMVLLGKIFQKDSSRSKLFL |
| UniProt | Q923Y8 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | N/A |
| 3D structure model | No available structures or models |
| BioLiP | N/A |
| Therapeutic Target Database | N/A |
| ChEMBL | CHEMBL4908 |
| IUPHAR | 364 |
| DrugBank | N/A |
| Name | CHEMBL3641694 |
|---|---|
| Molecular formula | C14H18N4O |
| IUPAC name | 1-methyl-N-(4-morpholin-2-ylphenyl)pyrazol-3-amine |
| Molecular weight | 258.325 |
| Hydrogen bond acceptor | 4 |
| Hydrogen bond donor | 2 |
| XlogP | 1.1 |
| Synonyms | BDBM129498 SCHEMBL12610056 US8802673, 140 |
| Inchi Key | CPAWBSUVPJCERW-UHFFFAOYSA-N |
| Inchi ID | InChI=1S/C14H18N4O/c1-18-8-6-14(17-18)16-12-4-2-11(3-5-12)13-10-15-7-9-19-13/h2-6,8,13,15H,7,9-10H2,1H3,(H,16,17) |
| PubChem CID | 68325752 |
| ChEMBL | CHEMBL3641694 |
| IUPHAR | N/A |
| BindingDB | 129498 |
| DrugBank | N/A |
Structure | 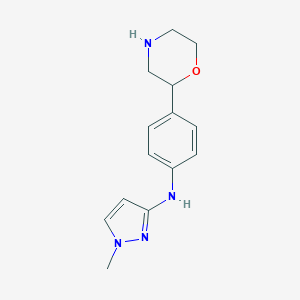 |
| Lipinski's druglikeness | This ligand satisfies Lipinski's rule of five. |
| Parameter | Value | Reference | Database source |
|---|---|---|---|
| Ki | 38.6 nM | , None | BindingDB,ChEMBL |
zhanglab![]() zhanggroup.org | +65-6601-1241 | Computing 1, 13 Computing Drive, Singapore 117417
zhanggroup.org | +65-6601-1241 | Computing 1, 13 Computing Drive, Singapore 117417