

You can:
| Name | Metabotropic glutamate receptor 2 |
|---|---|
| Species | Homo sapiens (Human) |
| Gene | GRM2 |
| Synonym | mGluR2 mGlu2 receptor metabotropic glutamate receptor 2 GPRC1B glutamate receptor |
| Disease | Central nervous system disease Anxiety disorder Bipolar disorder Major depressive disorder Mood disorder [ Show all ] |
| Length | 872 |
| Amino acid sequence | MGSLLALLALLLLWGAVAEGPAKKVLTLEGDLVLGGLFPVHQKGGPAEDCGPVNEHRGIQRLEAMLFALDRINRDPHLLPGVRLGAHILDSCSKDTHALEQALDFVRASLSRGADGSRHICPDGSYATHGDAPTAITGVIGGSYSDVSIQVANLLRLFQIPQISYASTSAKLSDKSRYDYFARTVPPDFFQAKAMAEILRFFNWTYVSTVASEGDYGETGIEAFELEARARNICVATSEKVGRAMSRAAFEGVVRALLQKPSARVAVLFTRSEDARELLAASQRLNASFTWVASDGWGALESVVAGSEGAAEGAITIELASYPISDFASYFQSLDPWNNSRNPWFREFWEQRFRCSFRQRDCAAHSLRAVPFEQESKIMFVVNAVYAMAHALHNMHRALCPNTTRLCDAMRPVNGRRLYKDFVLNVKFDAPFRPADTHNEVRFDRFGDGIGRYNIFTYLRAGSGRYRYQKVGYWAEGLTLDTSLIPWASPSAGPLPASRCSEPCLQNEVKSVQPGEVCCWLCIPCQPYEYRLDEFTCADCGLGYWPNASLTGCFELPQEYIRWGDAWAVGPVTIACLGALATLFVLGVFVRHNATPVVKASGRELCYILLGGVFLCYCMTFIFIAKPSTAVCTLRRLGLGTAFSVCYSALLTKTNRIARIFGGAREGAQRPRFISPASQVAICLALISGQLLIVVAWLVVEAPGTGKETAPERREVVTLRCNHRDASMLGSLAYNVLLIALCTLYAFKTRKCPENFNEAKFIGFTMYTTCIIWLAFLPIFYVTSSDYRVQTTTMCVSVSLSGSVVLGCLFAPKLHIILFQPQKNVVSHRAPTSRFGSAAARASSSLGQGSGSQFVPTVCNGREVVDSTTSSL |
| UniProt | Q14416 |
| Protein Data Bank | 5cnj, 5kzn, 5kzq, 5cni, 4xas, 4xaq |
| GPCR-HGmod model | N/A |
| 3D structure model | This structure is from PDB ID 5cnj. |
| BioLiP | BL0365442, BL0324559,BL0324560, BL0324557,BL0324558, BL0306697,BL0306698, BL0306695,BL0306696, BL0365443 |
| Therapeutic Target Database | T62820 |
| ChEMBL | CHEMBL5137 |
| IUPHAR | 290 |
| DrugBank | N/A |
| Name | (1S,3R)-ACPD |
|---|---|
| Molecular formula | C7H11NO4 |
| IUPAC name | (1S,3R)-1-aminocyclopentane-1,3-dicarboxylic acid |
| Molecular weight | 173.168 |
| Hydrogen bond acceptor | 5 |
| Hydrogen bond donor | 3 |
| XlogP | -3.2 |
| Synonyms | trans-( inverted question mark)-ACPD 1-Amino-1,3-dicarboxycyclopentane YFYNOWXBIBKGHB-FBCQKBJTSA- 1-Aminocyclopentane-1S,3R-dicarboxylic acid B6220 [ Show all ] |
| Inchi Key | YFYNOWXBIBKGHB-FBCQKBJTSA-N |
| Inchi ID | InChI=1S/C7H11NO4/c8-7(6(11)12)2-1-4(3-7)5(9)10/h4H,1-3,8H2,(H,9,10)(H,11,12)/t4-,7+/m1/s1 |
| PubChem CID | 104766 |
| ChEMBL | CHEMBL34453 |
| IUPHAR | 1365 |
| BindingDB | 66976 |
| DrugBank | N/A |
Structure | 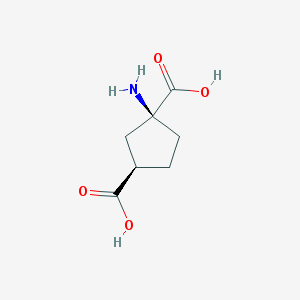 |
| Lipinski's druglikeness | This ligand satisfies Lipinski's rule of five. |
| Parameter | Value | Reference | Database source |
|---|---|---|---|
| Activity | 12000.0 nM | PMID12166935 | ChEMBL |
| EC50 | 2000.0 nM | PMID9572889, PMID12825948 | BindingDB,ChEMBL |
| EC50 | 5000.0 nM | PMID7738999 | BindingDB |
| EC50 | 7700.0 nM | PMID7738999 | BindingDB |
| EC50 | 12100.0 nM | PMID7738999 | BindingDB,ChEMBL |
| EC50 | 18000.0 nM | PMID9301676 | BindingDB,ChEMBL |
| Ki | 5000.0 nM | PMID10893301 | BindingDB,ChEMBL |
| Ki | 63095.7 nM | PMID10530814 | IUPHAR |
zhanglab![]() zhanggroup.org | +65-6601-1241 | Computing 1, 13 Computing Drive, Singapore 117417
zhanggroup.org | +65-6601-1241 | Computing 1, 13 Computing Drive, Singapore 117417