

You can:
| Name | Metabotropic glutamate receptor 5 |
|---|---|
| Species | Homo sapiens (Human) |
| Gene | GRM5 |
| Synonym | mGlu5 receptor glutamate receptor GPRC1E mGluR5 |
| Disease | Central nervous system disease Chronic neuropathic pain Fragile X syndrome Major depressive disorder; GERD; Chronic neuropathic pain Autism [ Show all ] |
| Length | 1212 |
| Amino acid sequence | MVLLLILSVLLLKEDVRGSAQSSERRVVAHMPGDIIIGALFSVHHQPTVDKVHERKCGAVREQYGIQRVEAMLHTLERINSDPTLLPNITLGCEIRDSCWHSAVALEQSIEFIRDSLISSEEEEGLVRCVDGSSSSFRSKKPIVGVIGPGSSSVAIQVQNLLQLFNIPQIAYSATSMDLSDKTLFKYFMRVVPSDAQQARAMVDIVKRYNWTYVSAVHTEGNYGESGMEAFKDMSAKEGICIAHSYKIYSNAGEQSFDKLLKKLTSHLPKARVVACFCEGMTVRGLLMAMRRLGLAGEFLLLGSDGWADRYDVTDGYQREAVGGITIKLQSPDVKWFDDYYLKLRPETNHRNPWFQEFWQHRFQCRLEGFPQENSKYNKTCNSSLTLKTHHVQDSKMGFVINAIYSMAYGLHNMQMSLCPGYAGLCDAMKPIDGRKLLESLMKTNFTGVSGDTILFDENGDSPGRYEIMNFKEMGKDYFDYINVGSWDNGELKMDDDEVWSKKSNIIRSVCSEPCEKGQIKVIRKGEVSCCWTCTPCKENEYVFDEYTCKACQLGSWPTDDLTGCDLIPVQYLRWGDPEPIAAVVFACLGLLATLFVTVVFIIYRDTPVVKSSSRELCYIILAGICLGYLCTFCLIAKPKQIYCYLQRIGIGLSPAMSYSALVTKTNRIARILAGSKKKICTKKPRFMSACAQLVIAFILICIQLGIIVALFIMEPPDIMHDYPSIREVYLICNTTNLGVVTPLGYNGLLILSCTFYAFKTRNVPANFNEAKYIAFTMYTTCIIWLAFVPIYFGSNYKIITMCFSVSLSATVALGCMFVPKVYIILAKPERNVRSAFTTSTVVRMHVGDGKSSSAASRSSSLVNLWKRRGSSGETLRYKDRRLAQHKSEIECFTPKGSMGNGGRATMSSSNGKSVTWAQNEKSSRGQHLWQRLSIHINKKENPNQTAVIKPFPKSTESRGLGAGAGAGGSAGGVGATGGAGCAGAGPGGPESPDAGPKALYDVAEAEEHFPAPARPRSPSPISTLSHRAGSASRTDDDVPSLHSEPVARSSSSQGSLMEQISSVVTRFTANISELNSMMLSTAAPSPGVGAPLCSSYLIPKEIQLPTTMTTFAEIQPLPAIEVTGGAQPAAGAQAAGDAARESPAAGPEAAAAKPDLEELVALTPPSPFRDSVDSGSTTPNSPVSESALCIPSSPKYDTLIIRDYTQSSSSL |
| UniProt | P41594 |
| Protein Data Bank | 4oo9, 5cgc, 5cgd, 6ffh, 6ffi, 6n4x, 6n4y, 6n50, 6n51, 3lmk |
| GPCR-HGmod model | N/A |
| 3D structure model | This structure is from PDB ID 4oo9. |
| BioLiP | BL0176927,BL0176931, BL0176928,BL0176929,BL0176930, BL0281199, BL0322076, BL0322077, BL0407724, BL0438693, BL0438694, BL0438695,BL0438696,BL0438697, BL0407725, BL0438698,BL0438699 |
| Therapeutic Target Database | T99347 |
| ChEMBL | CHEMBL3227 |
| IUPHAR | 293 |
| DrugBank | BE0001192 |
| Name | SCHEMBL11235794 |
|---|---|
| Molecular formula | C5H9NO4 |
| IUPAC name | (2S)-2-amino-5-hydroxy-5-oxopentanoate;hydron |
| Molecular weight | 147.13 |
| Hydrogen bond acceptor | 5 |
| Hydrogen bond donor | 3 |
| XlogP | None |
| Synonyms | N/A |
| Inchi Key | WHUUTDBJXJRKMK-VKHMYHEASA-N |
| Inchi ID | InChI=1S/C5H9NO4/c6-3(5(9)10)1-2-4(7)8/h3H,1-2,6H2,(H,7,8)(H,9,10)/t3-/m0/s1 |
| PubChem CID | 88747398 |
| ChEMBL | CHEMBL575060 |
| IUPHAR | 1369 |
| BindingDB | 17657 |
| DrugBank | DB00142 |
Structure | 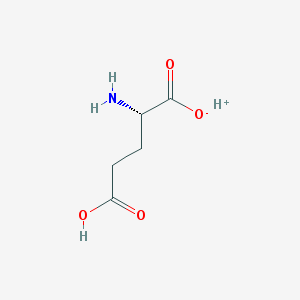 |
| Lipinski's druglikeness | Partition coefficient log P of this ligand is not available. |
| Parameter | Value | Reference | Database source |
|---|---|---|---|
| Activity | 4000.0 nM | PMID12166935 | ChEMBL |
| EC50 | 3100.0 nM | PMID9572889 | ChEMBL |
| EC50 | 3162.28 - 10000.0 nM | PMID10443583 | IUPHAR |
| EC50 | 4100.0 nM | PMID7738999 | BindingDB,ChEMBL |
| EC50 | 4700.0 nM | MedChemComm, (2011) 2:11:1120 | ChEMBL |
| EC50 | 631000.0 nM | PMID19819046 | ChEMBL |
| Ki | 390.0 nM | PMID17725337 | BindingDB,ChEMBL |
| Ki | 1160.0 nM | PMID19042134 | BindingDB,ChEMBL |
| Ki | 3000.0 nM | PMID9131252 | BindingDB |
| Ki | 4677.35 nM | MedChemComm, (2011) 2:11:1120 | ChEMBL |
| Ki | 11400.0 nM | PMID10893301 | BindingDB,ChEMBL |
zhanglab![]() zhanggroup.org | +65-6601-1241 | Computing 1, 13 Computing Drive, Singapore 117417
zhanggroup.org | +65-6601-1241 | Computing 1, 13 Computing Drive, Singapore 117417