

You can:
| Name | Muscarinic acetylcholine receptor M1 |
|---|---|
| Species | Rattus norvegicus (Rat) |
| Gene | Chrm1 |
| Synonym | cholinergic receptor, muscarinic 1 cholinergic receptor, muscarinic 1, CNS cholinergic receptor M1 muscarinic acetylcholine receptor M1 receptor [ Show all ] |
| Disease | N/A for non-human GPCRs |
| Length | 460 |
| Amino acid sequence | MNTSVPPAVSPNITVLAPGKGPWQVAFIGITTGLLSLATVTGNLLVLISFKVNTELKTVNNYFLLSLACADLIIGTFSMNLYTTYLLMGHWALGTLACDLWLALDYVASNASVMNLLLISFDRYFSVTRPLSYRAKRTPRRAALMIGLAWLVSFVLWAPAILFWQYLVGERTVLAGQCYIQFLSQPIITFGTAMAAFYLPVTVMCTLYWRIYRETENRARELAALQGSETPGKGGGSSSSSERSQPGAEGSPESPPGRCCRCCRAPRLLQAYSWKEEEEEDEGSMESLTSSEGEEPGSEVVIKMPMVDSEAQAPTKQPPKSSPNTVKRPTKKGRDRGGKGQKPRGKEQLAKRKTFSLVKEKKAARTLSAILLAFILTWTPYNIMVLVSTFCKDCVPETLWELGYWLCYVNSTVNPMCYALCNKAFRDTFRLLLLCRWDKRRWRKIPKRPGSVHRTPSRQC |
| UniProt | P08482 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | N/A |
| 3D structure model | No available structures or models |
| BioLiP | N/A |
| Therapeutic Target Database | N/A |
| ChEMBL | CHEMBL276 |
| IUPHAR | 13 |
| DrugBank | N/A |
| Name | pirenzepine |
|---|---|
| Molecular formula | C19H21N5O2 |
| IUPAC name | 11-[2-(4-methylpiperazin-1-yl)acetyl]-5H-pyrido[2,3-b][1,4]benzodiazepin-6-one |
| Molecular weight | 351.41 |
| Hydrogen bond acceptor | 5 |
| Hydrogen bond donor | 1 |
| XlogP | 0.1 |
| Synonyms | PDSP2_000949 AB00053603 pirenzepinedihydrochloride BPBio1_000196 RMHMFHUVIITRHF-UHFFFAOYSA-N [ Show all ] |
| Inchi Key | RMHMFHUVIITRHF-UHFFFAOYSA-N |
| Inchi ID | InChI=1S/C19H21N5O2/c1-22-9-11-23(12-10-22)13-17(25)24-16-7-3-2-5-14(16)19(26)21-15-6-4-8-20-18(15)24/h2-8H,9-13H2,1H3,(H,21,26) |
| PubChem CID | 4848 |
| ChEMBL | CHEMBL9967 |
| IUPHAR | 328 |
| BindingDB | 39341 |
| DrugBank | DB00670 |
Structure | 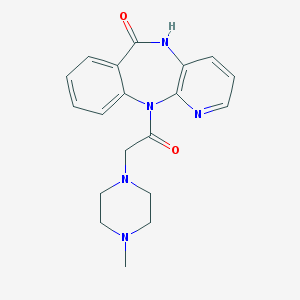 |
| Lipinski's druglikeness | This ligand satisfies Lipinski's rule of five. |
| Parameter | Value | Reference | Database source |
|---|---|---|---|
| IC50 | 3.8 nM | PMID10571170 | BindingDB,ChEMBL |
| IC50 | 120.0 nM | PMID3373484 | BindingDB,ChEMBL |
| Ki | <10000.0 nM | PMID3272174 | BindingDB |
| Ki | 5.2 nM | , Bioorg. Med. Chem. Lett., (1997) 7:8:979, PMID8464039, PMID1920350 | BindingDB,ChEMBL |
| Ki | 5.21 nM | PMID8464039, PMID1920350 | ChEMBL |
| Ki | 6.457 nM | PMID7932564, Bioorg. Med. Chem. Lett., (1995) 5:20:2325, PMID8246244, Bioorg. Med. Chem. Lett., (1995) 5:8:785 | ChEMBL |
| Ki | 6.46 nM | PMID8246244 | BindingDB |
| Ki | 6.5 nM | N/A | BindingDB |
| Ki | 6.99 nM | PMID2547939 | BindingDB |
| Ki | 9.0 nM | PMID1646322 | BindingDB |
| Ki | 9.55 nM | PMID2250662 | BindingDB |
| Ki | 12.5893 - 15.8489 nM | PMID1325587, PMID2704370 | IUPHAR |
| Ki | 12.59 nM | PMID2188581 | BindingDB |
| Ki | 16.0 nM | PMID2704370 | BindingDB |
| Ki | 23.4 nM | PMID1310135 | BindingDB |
| Ki | 29.0 nM | PMID2066986 | BindingDB,ChEMBL |
| Ki | 1.25893e+17 nM | PMID9703467 | ChEMBL |
| Potency | 100.0 nM | PubChem BioAssay data set | ChEMBL |
| Potency | 125.9 nM | PubChem BioAssay data set | ChEMBL |
zhanglab![]() zhanggroup.org | +65-6601-1241 | Computing 1, 13 Computing Drive, Singapore 117417
zhanggroup.org | +65-6601-1241 | Computing 1, 13 Computing Drive, Singapore 117417