

You can:
| Name | C-C chemokine receptor type 1 |
|---|---|
| Species | Homo sapiens (Human) |
| Gene | CCR1 |
| Synonym | macrophage inflammatory protein-1 alpha receptor MIP-1alpha-R MIP-1alpha/RANTES MIP-1alphaR MIP1aR [ Show all ] |
| Disease | Autoimmune diabetes Rheumatoid arthritis Chronic obstructive pulmonary disease |
| Length | 355 |
| Amino acid sequence | METPNTTEDYDTTTEFDYGDATPCQKVNERAFGAQLLPPLYSLVFVIGLVGNILVVLVLVQYKRLKNMTSIYLLNLAISDLLFLFTLPFWIDYKLKDDWVFGDAMCKILSGFYYTGLYSEIFFIILLTIDRYLAIVHAVFALRARTVTFGVITSIIIWALAILASMPGLYFSKTQWEFTHHTCSLHFPHESLREWKLFQALKLNLFGLVLPLLVMIICYTGIIKILLRRPNEKKSKAVRLIFVIMIIFFLFWTPYNLTILISVFQDFLFTHECEQSRHLDLAVQVTEVIAYTHCCVNPVIYAFVGERFRKYLRQLFHRRVAVHLVKWLPFLSVDRLERVSSTSPSTGEHELSAGF |
| UniProt | P32246 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | P32246 |
| 3D structure model | This predicted structure model is from GPCR-EXP P32246. |
| BioLiP | N/A |
| Therapeutic Target Database | T16016 |
| ChEMBL | CHEMBL2413 |
| IUPHAR | 58 |
| DrugBank | N/A |
| Name | CHEMBL3775436 |
|---|---|
| Molecular formula | C15H26ClNO3 |
| IUPAC name | (2R)-2-[2-[2-(2,6-dimethylphenoxy)ethoxy]ethylamino]propan-1-ol;hydrochloride |
| Molecular weight | 303.827 |
| Hydrogen bond acceptor | 4 |
| Hydrogen bond donor | 3 |
| XlogP | None |
| Synonyms | SCHEMBL16446074 |
| Inchi Key | BFBVSDYKRDBSFJ-PFEQFJNWSA-N |
| Inchi ID | InChI=1S/C15H25NO3.ClH/c1-12-5-4-6-13(2)15(12)19-10-9-18-8-7-16-14(3)11-17;/h4-6,14,16-17H,7-11H2,1-3H3;1H/t14-;/m1./s1 |
| PubChem CID | 117858220 |
| ChEMBL | CHEMBL3775436 |
| IUPHAR | N/A |
| BindingDB | N/A |
| DrugBank | N/A |
Structure | 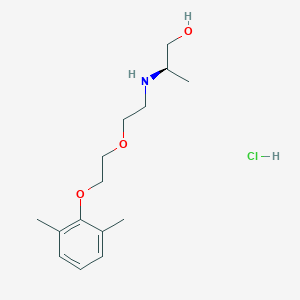 |
| Lipinski's druglikeness | Partition coefficient log P of this ligand is not available. |
| Parameter | Value | Reference | Database source |
|---|---|---|---|
| Inhibition | -11.2 % | PMID26988801 | ChEMBL |
zhanglab![]() zhanggroup.org | +65-6601-1241 | Computing 1, 13 Computing Drive, Singapore 117417
zhanggroup.org | +65-6601-1241 | Computing 1, 13 Computing Drive, Singapore 117417