

You can:
| Name | 5-hydroxytryptamine receptor 2B |
|---|---|
| Species | Homo sapiens (Human) |
| Gene | HTR2B |
| Synonym | stomach fundus serotonin receptor serotonin receptor 2B 5-HT2F 5-HT-2B 5-HT2B [ Show all ] |
| Disease | Psychoses Migraine Irritable bowel syndrome Depression; Cerebral infarction Coronary heart disease [ Show all ] |
| Length | 481 |
| Amino acid sequence | MALSYRVSELQSTIPEHILQSTFVHVISSNWSGLQTESIPEEMKQIVEEQGNKLHWAALLILMVIIPTIGGNTLVILAVSLEKKLQYATNYFLMSLAVADLLVGLFVMPIALLTIMFEAMWPLPLVLCPAWLFLDVLFSTASIMHLCAISVDRYIAIKKPIQANQYNSRATAFIKITVVWLISIGIAIPVPIKGIETDVDNPNNITCVLTKERFGDFMLFGSLAAFFTPLAIMIVTYFLTIHALQKKAYLVKNKPPQRLTWLTVSTVFQRDETPCSSPEKVAMLDGSRKDKALPNSGDETLMRRTSTIGKKSVQTISNEQRASKVLGIVFFLFLLMWCPFFITNITLVLCDSCNQTTLQMLLEIFVWIGYVSSGVNPLVYTLFNKTFRDAFGRYITCNYRATKSVKTLRKRSSKIYFRNPMAENSKFFKKHGIRNGINPAMYQSPMRLRSSTIQSSSIILLDTLLLTENEGDKTEEQVSYV |
| UniProt | P41595 |
| Protein Data Bank | 5tud, 5tvn, 6drx, 4ib4, 4nc3, 6dry, 6drz, 6ds0 |
| GPCR-HGmod model | P41595 |
| 3D structure model | This structure is from PDB ID 5tud. |
| BioLiP | BL0425280, BL0425283, BL0425282, BL0239858, BL0239859, BL0265182, BL0265183, BL0368464, BL0425281, BL0385686,BL0385687, BL0425284, BL0425285, BL0425286, BL0368465, BL0425287 |
| Therapeutic Target Database | T31204 |
| ChEMBL | CHEMBL1833 |
| IUPHAR | 7 |
| DrugBank | BE0000393 |
| Name | ritanserin |
|---|---|
| Molecular formula | C27H25F2N3OS |
| IUPAC name | 6-[2-[4-[bis(4-fluorophenyl)methylidene]piperidin-1-yl]ethyl]-7-methyl-[1,3]thiazolo[3,2-a]pyrimidin-5-one |
| Molecular weight | 477.574 |
| Hydrogen bond acceptor | 6 |
| Hydrogen bond donor | 0 |
| XlogP | 5.2 |
| Synonyms | Tox21_110251 CHEMBL267777 SBI-0051053.P003 DivK1c_000192 FT-0630948 [ Show all ] |
| Inchi Key | JUQLTPCYUFPYKE-UHFFFAOYSA-N |
| Inchi ID | InChI=1S/C27H25F2N3OS/c1-18-24(26(33)32-16-17-34-27(32)30-18)12-15-31-13-10-21(11-14-31)25(19-2-6-22(28)7-3-19)20-4-8-23(29)9-5-20/h2-9,16-17H,10-15H2,1H3 |
| PubChem CID | 5074 |
| ChEMBL | CHEMBL267777 |
| IUPHAR | 97 |
| BindingDB | 50001775 |
| DrugBank | DB12693 |
Structure | 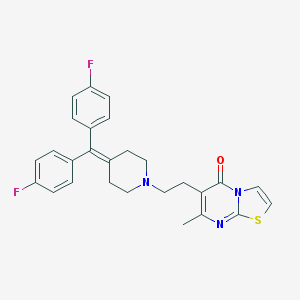 |
| Lipinski's druglikeness | This ligand has a partition coefficient log P greater than 5. |
| Parameter | Value | Reference | Database source |
|---|---|---|---|
| IC50 | 3.0 nM | PMID16469866 | BindingDB |
| Ki | 0.4 nM | PMID9459568 | PDSP,BindingDB |
| Ki | 0.630957 - 1.99526 nM | PMID15322733, PMID10188965, PMID9459568, PMID10455251 | IUPHAR |
| Ki | 1.0 nM | PMID10455251 | PDSP,BindingDB |
| Ki | 1.25893 nM | PMID11882920 | PDSP |
| Ki | 1.26 nM | PMID11882920 | BindingDB |
| Ki | 1.64 nM | PMID9459568 | PDSP,BindingDB |
| Ki | 1.73 nM | PMID8632342 | PDSP,BindingDB |
| Ki | 2.13796 nM | PMID15322733 | PDSP |
| Ki | 2.14 nM | PMID15322733 | BindingDB |
| Ki | 5.01 nM | PMID7582481, PMID10821800 | PDSP,BindingDB |
| Ki | 5.01187 nM | PMID10821800 | PDSP |
| Ki | 6.45 nM | PMID9225287 | PDSP,BindingDB |
zhanglab![]() zhanggroup.org | +65-6601-1241 | Computing 1, 13 Computing Drive, Singapore 117417
zhanggroup.org | +65-6601-1241 | Computing 1, 13 Computing Drive, Singapore 117417