UniBioMap is an AI-ready, updatable biomedical knowledge graph centered on proteins and small molecules. It harmonizes identifiers across six major entity types and integrates evidence from dozens of public resources into a unified, provenance-aware graph. An AI-based confident-learning pipeline further assigns reliability scores and expands coverage with high-confidence predictions. This page summarizes UniBioMap's methodology, key statistics, and how to use the web server and datasets.
- Overview
UniBioMap addresses data fragmentation by normalizing entities (proteins, compounds, diseases, pathways, GO, and phenotypes) and unifying relations into a consistent, machine-learning–friendly schema. The platform couples broad, curated integration with a confidence-aware recommendation layer, enabling prioritization of reliable edges and hypothesis generation through predicted associations.

Figure 1. Overview of the UniBioMap workflow and system architecture.
- System Architecture & Data Integration
UniBioMap employs a modular, automated pipeline for data ingestion, identifier normalization, relation unification, and graph assembly. Proteins are normalized to UniProt accessions; compounds are unified by chemical structure identifiers; diseases, pathways, and other entities are mapped through established hubs to ensure cross-database consistency. The result is a clean core graph for modeling, complemented by an auxiliary property layer that preserves rich biological context and provenance.

Figure 2. Landscape of UniBioMap: entity coverage, relation composition, and comparisons with existing biomedical KGs.
- Confident Learning & Knowledge Graph Completion
To prioritize reliability and expand coverage, UniBioMap integrates a confident-learning workflow: out-of-fold predictions from KGC models are calibrated to estimate edge confidence, noisy links are down-weighted, and a refined model proposes additional high-confidence relations. This iterative predict–estimate–refine cycle yields a KG with improved fidelity and extensive, ranked hypotheses around protein- and compound-centered biology.
- Web Server & Usage
The UniBioMap web server provides two primary modules:
- Search system. Retrieve entities by internal IDs or external identifiers (e.g., UniProt, UniChem) or by names. Compound pages support structure-based retrieval (similarity and substructure) via a built-in sketcher and SMILES input. Entity pages aggregate metadata, ontology annotations, and evidence-linked relations, with an instant local subgraph preview.
- Query system. Select up to five entities across supported types to generate multi-hop subgraphs via sampling. Results are shown in synchronized visualization and table views, with Graph, Sankey, and Statistics modes for interactive analysis and export.
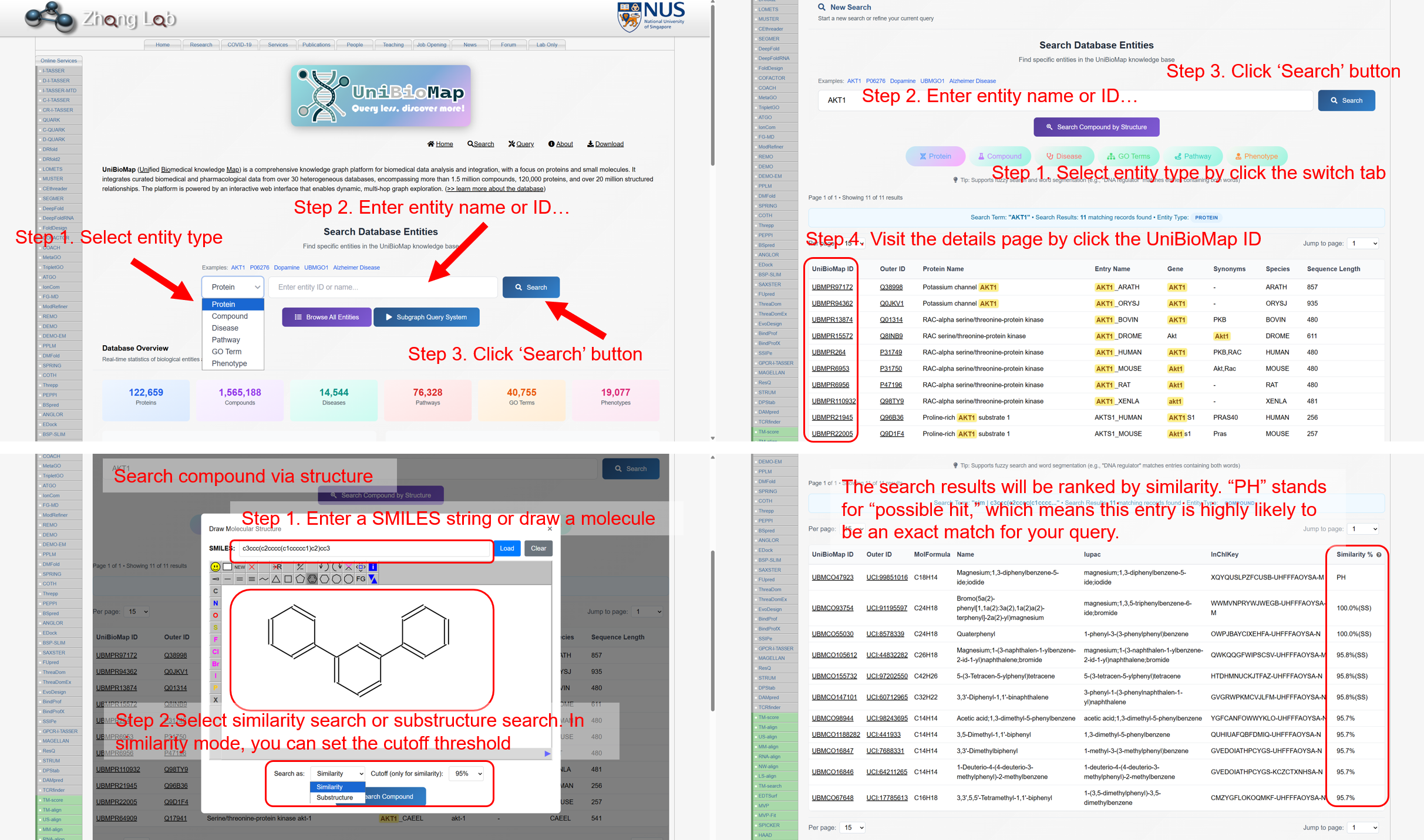
Figure 3A. Search interface and entity retrieval.
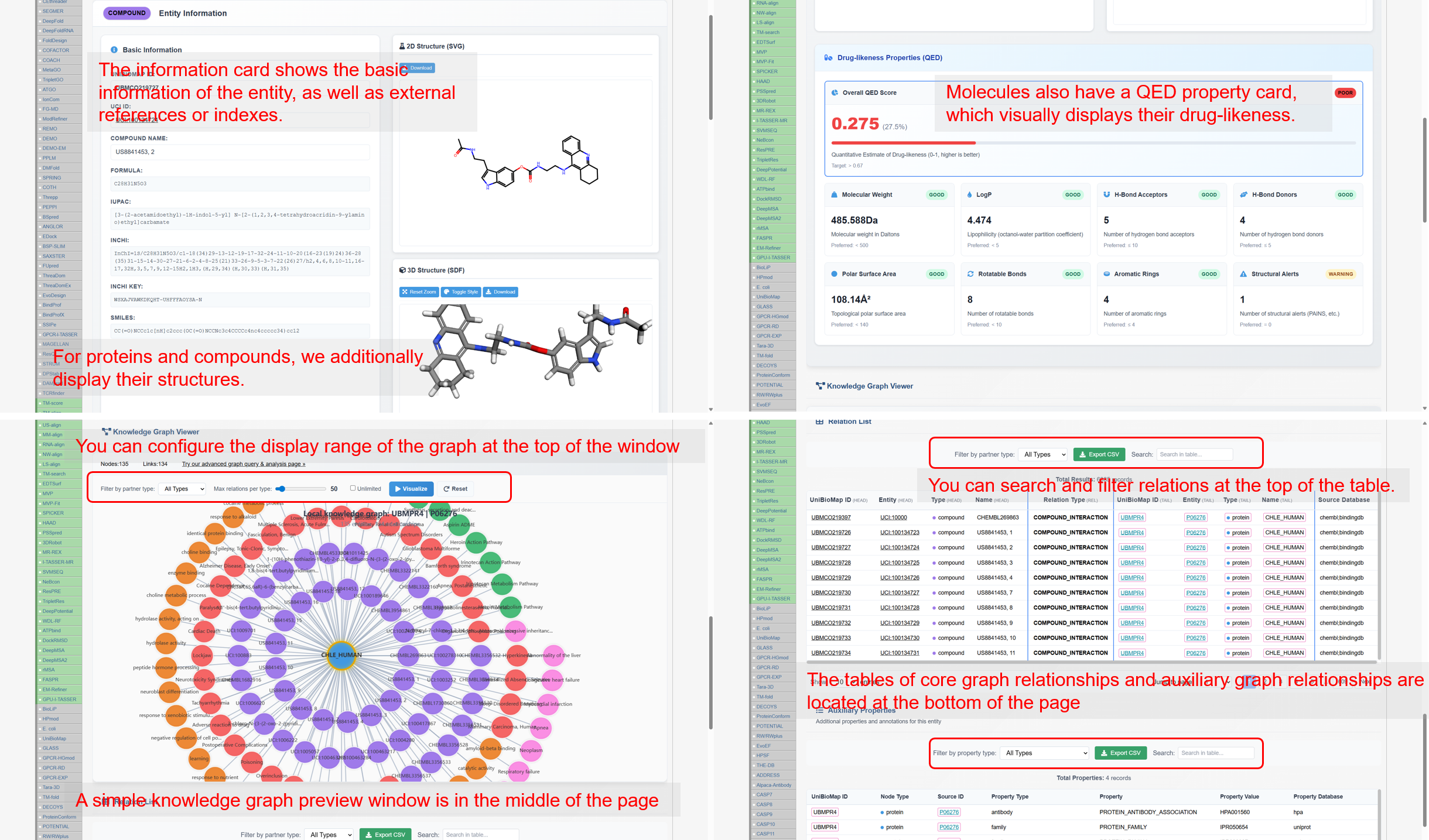
Figure 3B. Entity detail page with integrated metadata, structure viewers, and local subgraph.
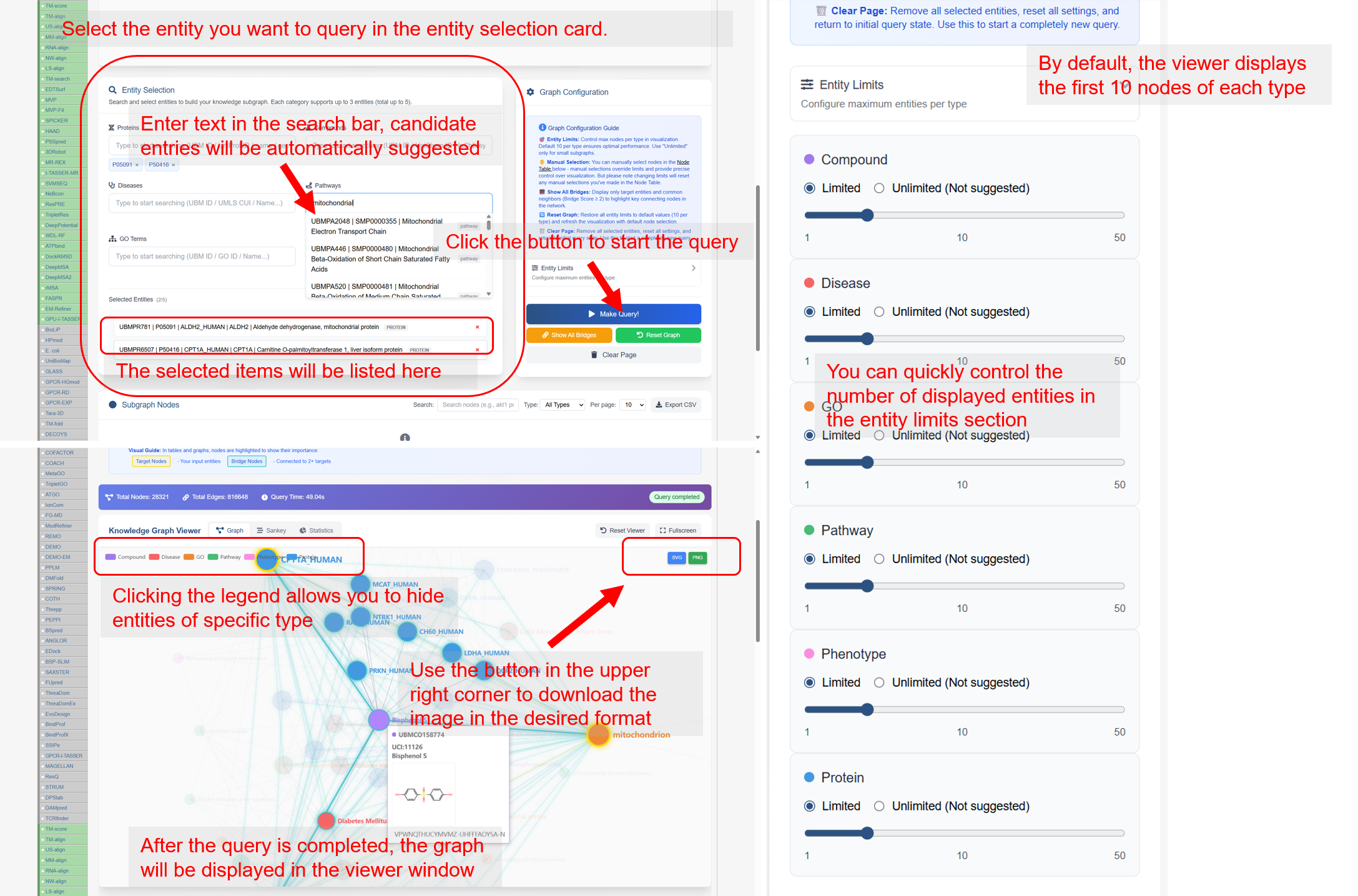
Figure 3C. Query interface for multi-entity subgraph construction.
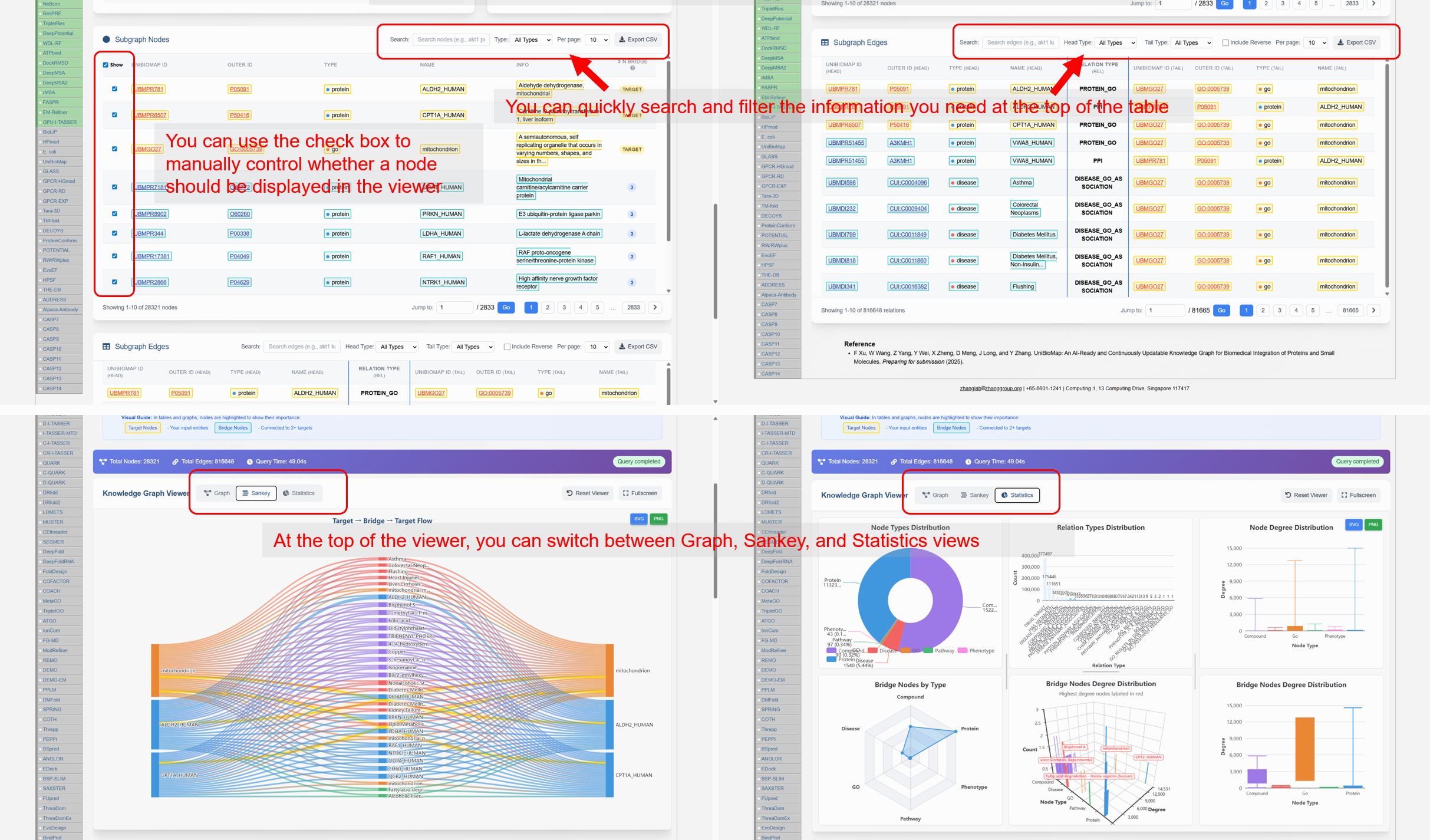
Figure 3D. Visualization modes (Graph, Sankey, Statistics) and synchronized tables.



