

| 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 | |
| | | | | | | | | | | | | | | | | |
| MEKINNVTEFIFWGLSQSPEIEKVCFVVFSFFYIIILLGNLLIMLTVCLSNLFKSPMYFFLSFLSFVDICYSSVTAPKMIVDLLAKDKTISYVGCMLQLFGVHFFGCTEIFILTVMAYDRYVAICKPLHYMTIMNRETCNKMLLGTWVGGFLHSIIQVALVVQLPFCGPNEIDHYFCDVHPVLKLACTETYIVGVVVTANSGTIALGSFVILLISYSIILVSLRKQSAEGRRKALSTCGSHIAMVVIFFGPCTFMYMRPDTTFSEDKMVAVFYTIITPMLNPLIYTLRNAEVKNAMKKLWGRNVFLEAKGK | |
| CCCCCCCCSSSSSCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHSSSSSCCCCCCCCHHHHHHHHHHHHHHHHCCCCHHHHHHHHCCCCSSCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCHHCCCHHHHHHHHHHHHHHHHHHHHHHHHHHSSCCCCCCCCCCCSSCCCHHHHHHHHCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCHHHHHHHHHHHHHHHHHHHHHHCCSSSSSCCCCCCCCCCCSSSHHHHCHHHHCCCHHHCCCCHHHHHHHHHHHHHHCCCCCCCC | |
| 99997411468816999821589999999999999999788745523217999997899988699886502146748999987339967858999999999999999899999999865178626110260026776999999999999999999999992408988988267664284887888422305344225667659999999999999999999960137402018998898799942045106546886889999874245023434033010463334629999999999851010003589 |
| 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 | |
| | | | | | | | | | | | | | | | | |
| MEKINNVTEFIFWGLSQSPEIEKVCFVVFSFFYIIILLGNLLIMLTVCLSNLFKSPMYFFLSFLSFVDICYSSVTAPKMIVDLLAKDKTISYVGCMLQLFGVHFFGCTEIFILTVMAYDRYVAICKPLHYMTIMNRETCNKMLLGTWVGGFLHSIIQVALVVQLPFCGPNEIDHYFCDVHPVLKLACTETYIVGVVVTANSGTIALGSFVILLISYSIILVSLRKQSAEGRRKALSTCGSHIAMVVIFFGPCTFMYMRPDTTFSEDKMVAVFYTIITPMLNPLIYTLRNAEVKNAMKKLWGRNVFLEAKGK | |
| 87662200000000004245001000031333233233323300000200430300001002300120011000000200000025321000500000001103311310100020030000000221101000033100000120131023013001310230300011220001002210030000202300030022013202310331331121002001321461222011003002100301321010000003331330100003102310330130000214402300230134303246668 |
| Rank | PDB Hit | Iden1 | Iden2 | Cov. | Norm. Z-score | Download Align. | 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 | | | | | | | | | | | | | | | | |||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sec.Str Seq | CCCCCCCCSSSSSCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHSSSSSCCCCCCCCHHHHHHHHHHHHHHHHCCCCHHHHHHHHCCCCSSCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCHHCCCHHHHHHHHHHHHHHHHHHHHHHHHHHSSCCCCCCCCCCCSSCCCHHHHHHHHCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCHHHHHHHHHHHHHHHHHHHHHHCCSSSSSCCCCCCCCCCCSSSHHHHCHHHHCCCHHHCCCCHHHHHHHHHHHHHHCCCCCCCC MEKINNVTEFIFWGLSQSPEIEKVCFVVFSFFYIIILLGNLLIMLTVCLSNLFKSPMYFFLSFLSFVDICYSSVTAPKMIVDLLAKDKTISYVGCMLQLFGVHFFGCTEIFILTVMAYDRYVAICKPLHYMTIMNRETCNKMLLGTWVGGFLHSIIQVALVVQLPFCGPNEIDHYFCDVHPVLKLACTETYIVGVVVTANSGTIALGSFVILLISYSIILVSLRKQSAEGRRKALSTCGSHIAMVVIFFGPCTFMYMRPDTTFSEDKMVAVFYTIITPMLNPLIYTLRNAEVKNAMKKLWGRNVFLEAKGK | |||||||||||||||||||||||||
| 1 | 3emlA | 0.21 | 0.22 | 0.88 | 3.41 | Download | ------------------IMGSSVYITVELAIAVLAILGNVLVCWAVWLNSNLQNVTNYFVVSLAAADIAVGVLAIPFAITIS---GFCAACHGCLFIACFVLVLTQSSIFSLLAIAIDRYIAIRIPLRYNGLVTGTRAKGIIAICWVLSFAIGLTPMLGWNNCGGCGEGQVACLFEDVVPMNYMVYF-----------NFFACVLVPLLLMLGVYLRIFLAARQLVHAAKSLAIIVGLFALCWLPLHII-NCFTFFCPDCSHAWLMYLAIVLSHTNSVVNPFIYAYRIREFRQTFRKIIRSHVLRQ---- | |||||||||||||||||||
| 2 | 5tgzA | 0.19 | 0.22 | 0.89 | 2.18 | Download | -ENFMDIECF----MVLNPSQQLAIAVLSLTLGTFTVLENLLVLCVILHSRSLRCPSYHFIGSLAVADLLGSVIFVYSFIDFHVFHRK-DSRNVFLFKLGGVTASFTASVGSLFLAAIDRYISIHRPLAYKRIVTRPKAVVAFCLMWTIAIVIAVL--------PLLGWNCEK---------LQSVCSDIFHIDKTYLMFWIGVVSVLLLFIVYAYMYILWKAHSHAPDQARELAKTLVLILVVLIICWGPLLAIMVKMNKLIKTVFAFCSMLCLLNSTVNPIIYALRSKDLRHAFRSM------------ | |||||||||||||||||||
| 3 | 5tgzA | 0.17 | 0.22 | 0.87 | 2.03 | Download | -GGRGENFMDIECFMVLNPSQQLAIAVLSLTLGTFTVLENLLVLCVILHSRSLRCPSYHFIGSLAVADLLGSVIFVYSFIDFH-VFHRKDSRNVFLFKLGGVTASFTASVGSLFLAAIDRYISIHRPLAYKRIVTRPKAVVAFCLMWTIAIVIAVLPLLGSDIFPHI--------------------------DKTYLMFWIGVVSVLLLFIVYAYMYILWKAHSHQARMDIELAKTLVLILVVLIICWGPLLAIMVGKMNLIKTVFAFCSMLCLLNSTVNPIIYALRSKDLRHAFRSMF----------- | |||||||||||||||||||
| 4 | 4ib4 | 0.19 | 0.24 | 0.88 | 1.52 | Download | --------------EE-QGNKLHWAALLILMVIIPTIGGNTLVILAVSLEKKLQYATNYFLMSLAVADLLVGLFVMPIALLTIMFAMWPLPLVLCPAWLFLDVLFSTASIWHLCAISVDRYIAIKKPIQANQYNSRATAFIKITVVWLISIGIAIPVPIKG--IE----TNPNNITCVLTK--------E-RFGDFMLFGSLAAFFTPLAIMIVTYFLTIHALQKKAADNEQRASKVLGIVFFLFLLMWCPFFITNITLVLCQQMLLEIFVWIGYVSSGVNPLVYTLFNKTFRDAFGRYITCNYR------ | |||||||||||||||||||
| 5 | 4yay | 0.16 | 0.23 | 0.90 | 1.21 | Download | KTRNAYIQKYLILNSSDCNYIFVMIPTLYSIIFVVGIFGNSLVVIVIYFYMKLKTVASVFLLNLALADLCFLLTLPLWAVYTAMEYRWPFGNYLCKIASASVSFNLYASVFLLTCLSIDRYLAIVHPMKSRLRRTMLVAKVTCIIIWLLAGLASLPAIIHRNVFF-------IITVCAFHYE--------TLPIGLGLTKNILGFLFPFLIILTSYTLIWKALK-----KNDDIFKIIMAIVLFFFFSWIPHQIFTFLDVLIVDTAMPITICIAYFNNCLNPLFYGFLGKKFKRYFLQLL----------- | |||||||||||||||||||
| 6 | 3emlA | 0.21 | 0.22 | 0.88 | 3.57 | Download | -------------------MGSSVYITVELAIAVLAILGNVLVCWAVWLNSNLQNVTNYFVVSLAAADIAVGVLAIPFAIT--ISTGFCAACHGCLFIACFVLVLTQSSIFSLLAIAIDRYIAIRIPLRYNGLVTGTRAKGIIAICWVLSFAIGLTPMLGWNNCGGCGEGQVACLFEDVVP-----------MNYMVYFNFFACVLVPLLLMLGVYLRIFLAARRQLVHAAKSLAIIVGLFALCWLPLHIINCFTFFCPDCSHAWLMYLAIVLSHTNSVVNPFIYAYRIREFRQTFRKIIRSHVLRQ---- | |||||||||||||||||||
| 7 | 4iaq | 0.20 | 0.23 | 0.85 | 1.73 | Download | ------------------LPWKVLLVMLLALITLATTLSNAFVIATVYRTRKLHTPANYLIASLAVTDLLVSILVMPISTMYTVTGRWTLGQVVCDFWLSSDITCCTASIWHLCVIALDRYWAITDAVEYSAKRTPKRAAVMIALVWVFSISISLPPFF-WRQASECVVNT-----------------DHI---LYTVYSTVGAFYFPTLLLIALYGRIYVEARSRIADRERKATKTLGIILGAFIVCWLPFFIISLVMPIH-LAIFDFFTWLGYLNSLINPIIYTMSNEDFKQAFHKLIRFK-------- | |||||||||||||||||||
| 8 | 2z73A | 0.16 | 0.19 | 0.96 | 2.81 | Download | WYNPSIVVHPHWREFDQVPAVYYSLGIFIGICGIIGCGGNGIVIYLFTKTKSLQTPANMFIINLAFSDFTFSLVNGPLMTISCFLKKWIFGFAACKVYGFIGGIFGFMSIMTMAMISIDRYNVIGRPMAASKKMSHRRAFIMIIFVWLWSVLWAIGPIFGTLEGVLC------NCSFDY-----ISRDSTTRSNILCMFILGF--FGPILIIFFCYFNIVMQAGANAEMRLAKISIVIVSQFLLSWSPYAVVALLAQFGPWVTPYAAQLPVMFAKASAIHNPMIYSVSHPKFREAISQTFPWVLTCCQFDD | |||||||||||||||||||
| 9 | 5tgzA | 0.18 | 0.22 | 0.89 | 4.80 | Download | GENFMDIECFMVL----NPSQQLAIAVLSLTLGTFTVLENLLVLCVILHSRSLRCPSYHFIGSLAVADLLGSVIFVYSFIDFHVFH-RKDSRNVFLFKLGGVTASFTASVGSLFLAAIDRYISIHRPLAYKRIVTRPKAVVAFCLMWTIAIVIAVLPLLG---------NCEK---------LQSVCSDIFHIDKTYLMFWIGVVSVLLLFIVYAYMYILWKAHSHAPDQAIELAKTLVLILVVLIICWGPLLAIMVYDVFGKMN-FAFCSMLCLLNSTVNPIIYALRSKDLRHAFRSMF----------- | |||||||||||||||||||
| 10 | 2ydoA | 0.18 | 0.21 | 0.93 | 5.63 | Download | ---------------------SSVYITVELAIAVLAILGNVLVCWAVWLNSNLQNVTNYFVVSAAAADILVGVLAIPFAIA--ISTGFCAACHGCLFIACFVLVLTASSIFSLLAIAIDRYIAIRIPLRYNGLVTGTRAKGIIAICWVLSFAIGLTPMLGWNNCGQPKEGKAHSQGCGEGQVACLFEDVVPMNYMVYFNFFACVLVPLLLMLGVYLRFLAARRQLKQMESTLQKEVHAAKSLAIIVGLFALCCFTFFCPDCSHAPLMYLAIVLSHTNSVVNPFIYAYRIREFRQTFRKIIRSHVLRQQEPF | |||||||||||||||||||
| ||||||||||||||||||||||||||
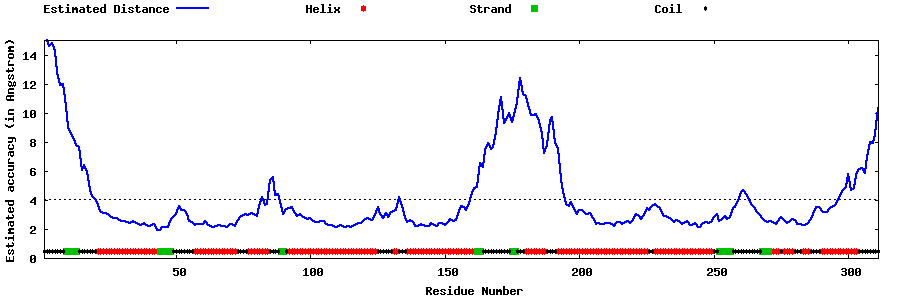
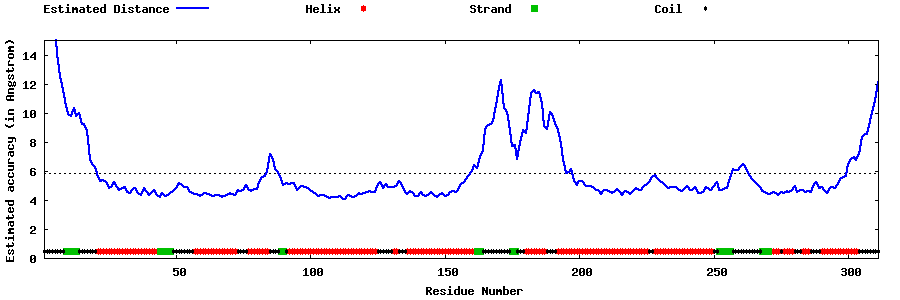
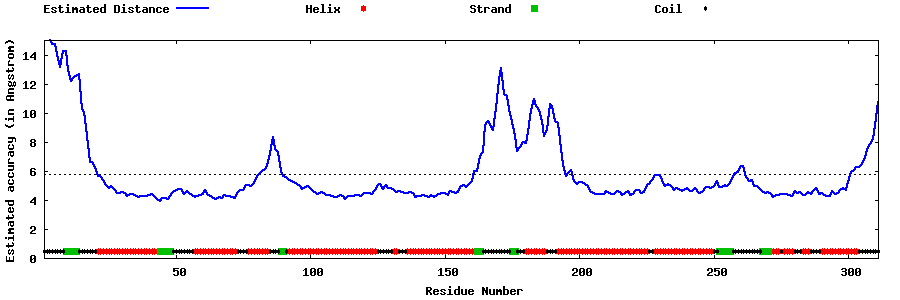
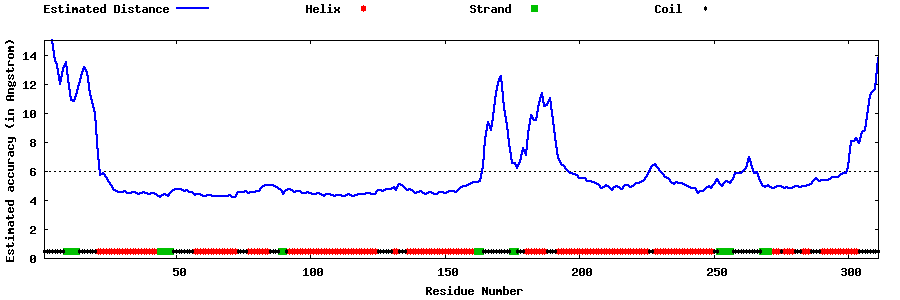
| Generated 3D models | Estimated local accuracy of models | ||
 | |||
|
|
|||
 | |||
|
| |||
 | |||
|
| |||
 | |||
|
| |||
 | |||
|
| |||