

| 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 | |
| | | | | | | | | | | | | | | | | |
| MTLGSLGNSSSSVSATFLLSGIPGLERMHIWISIPLCFMYLVSIPGNCTILFIIKTERSLHEPMYLFLSMLALIDLGLSLCTLPTVLGIFWVGAREISHDACFAQLFFIHCFSFLESSVLLSMAFDRFVAICHPLHYVSILTNTVIGRIGLVSLGRSVALIFPLPFMLKRFPYCGSPVLSHSYCLHQEVMKLACADMKANSIYGMFVIVSTVGIDSLLILFSYALILRTVLSIASRAERFKALNTCVSHICAVLLFYTPMIGLSVIHRFGKQAPHLVQVVMGFMYLLFPPVMNPIVYSVKTKQIRDRVTHAFCY | |
| CCCCCCCCCCCCCCHHHSSSCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCSSCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCCCCCHHHHHHHHHCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCHHHHHHHHHHHHCCCCCCCSSSSCCHHHHHHHHHHHCCC | |
| 96777789998766425047898986688999999999999999999999999956887532189999999999998998679999999966998578878999999999999999999999983416630566546222888999999999999999999889999964899999944762211265688716785375899999999999998999999999999998368887688898641499999999999989999746304788885799999999983886647822215479999999997269 |
| 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 | |
| | | | | | | | | | | | | | | | | |
| MTLGSLGNSSSSVSATFLLSGIPGLERMHIWISIPLCFMYLVSIPGNCTILFIIKTERSLHEPMYLFLSMLALIDLGLSLCTLPTVLGIFWVGAREISHDACFAQLFFIHCFSFLESSVLLSMAFDRFVAICHPLHYVSILTNTVIGRIGLVSLGRSVALIFPLPFMLKRFPYCGSPVLSHSYCLHQEVMKLACADMKANSIYGMFVIVSTVGIDSLLILFSYALILRTVLSIASRAERFKALNTCVSHICAVLLFYTPMIGLSVIHRFGKQAPHLVQVVMGFMYLLFPPVMNPIVYSVKTKQIRDRVTHAFCY | |
| 44221230324331320200000121320200011022022203321210000020151001000100000020010003002020000000214603140000001200310230110011002000000021010000003410010000003213011202101021031046200000001021002000230300111112003323333231032011100200010014601320120010000000003211321010010143101100010022100300330030000203300410031016 |
| Rank | PDB Hit | Iden1 | Iden2 | Cov. | Norm. Z-score | Download Align. | 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 | | | | | | | | | | | | | | | | |||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sec.Str Seq | CCCCCCCCCCCCCCHHHSSSCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCSSCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCCCCCHHHHHHHHHCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCHHHHHHHHHHHHCCCCCCCSSSSCCHHHHHHHHHHHCCC MTLGSLGNSSSSVSATFLLSGIPGLERMHIWISIPLCFMYLVSIPGNCTILFIIKTERSLHEPMYLFLSMLALIDLGLSLCTLPTVLGIFWVGAREISHDACFAQLFFIHCFSFLESSVLLSMAFDRFVAICHPLHYVSILTNTVIGRIGLVSLGRSVALIFPLPFMLKRFPYCGSPVLSHSYCLHQEVMKLACADMKANSIYGMFVIVSTVGIDSLLILFSYALILRTVLSIASRAERFKALNTCVSHICAVLLFYTPMIGLSVIHRFGKQAPHLVQVVMGFMYLLFPPVMNPIVYSVKTKQIRDRVTHAFCY | |||||||||||||||||||||||||
| 1 | 3emlA | 0.17 | 0.22 | 0.87 | 2.42 | Download | ------------------------IM-GSSVYITVELAIAVLAILGNVLVCWAVWLNSNLQNVTNYFVVSLAAADIAVGVLAIPFAITIST--GFCAACHGCLFIACFVLVLTQSSIFSLLAIAIDRYIAIRIPLRYNGLVTGTRAKGIIAICWVLSFAIGLTPMLGWNNCGGCGEGQVACL-----------FEDVVPMNYMVYFNFFACVLVPLLLMLGVYLRIFLAARRQLVHAAKSLAIIVGLFALCWLPLHIINCFTFFC---PDCSHAPLWLMYLAIVLSHTNSVVNPFIYAYRIREFRQTFRKIIRS | |||||||||||||||||||
| 2 | 5tgzA | 0.21 | 0.24 | 0.86 | 2.11 | Download | --------------------------SQQLAIAVLSLTLGTFTVLENLLVLCVILHSRSLRRPSYHFIGSLAVADLLGSVIFVYSFIDF-HVFHRKDSRNVFLFKLGGVTASFTASVGSLFLAAIDRYISIHRPLAYKRIVTRPKAVVAFCLMWTIAIVIAVLPLL-------GWNCEKLQSVC---------SDIFPHIDKTYLMFWIGVVSVLLLFIVYAYMYILWKAHSHAPDQARIELAKTLVLILVVLIICWGPLLAIMVYDVFGKMNKLIKTVFFCSMLCLLNSTVNPIIYALRSKDLRHAFRSMF-- | |||||||||||||||||||
| 3 | 4zwjA | 0.17 | 0.19 | 0.96 | 1.94 | Download | NFYVPFSNATGVVRSPFEYPQYYLAPWQFSMLAAYMFLLIVLGFPINFLTLYVTVQHKKLRTPLNYILLNLAVADLFMVLGGFTSTLYTSLHGYFVFGPTGCNLQGFFATLGGEIALWSLVVLAIERYVVVCKPM-SNFRFGENHAIMGVAFTWVMALACAAP--------PLAGWSRYLQCSCGID----YYTLKPEVNESFVIYMFVVHFTIPMIIIFFCYGQLVFTVKEAATQKAEKEVTRMVIIYVIAFLICWVPYASVAFYIFTHQGSCGPIFMTIPAFFAKSAAIYNPVIYIMMNKQFRNCMLTTICC | |||||||||||||||||||
| 4 | 3uon | 0.19 | 0.24 | 0.88 | 1.57 | Download | -------------------------TFEVVFIVLVAGSLSLVTIIGNILVMVSIKVNRHLQTVNNYFLFSLACADLIIGVFSMNLYTLYTVIGYWPLGPVVCDLWLALDYVVSNASVMNLLIISFDRYFCVTKPLTYPVKRTTKMAGMMIAAAWVLSFILWAPAILFWQFIV--GVRTVEDGECYIQFF---------SNAAVTFGTAIAAFYLPVIIMTVLYWHISRASKSRINISREKKVTRTILAILLAFIITWAPYNVMVLINTFCAPCIPNTVWTIGYWLCYINSTINPACYALCNATFKKTFKHLLM- | |||||||||||||||||||
| 5 | 3uonA | 0.19 | 0.21 | 0.87 | 1.14 | Download | -------------------------TFEVVFIVLVAGSLSLVTIIGNILVMVSIKVNRHLQTVNNYFLFSLACADLIIGVFSMNLYTLYTVIGYWPLGPVVCDLWLALDYVVSNASVMNLLIISFDRYFCVTKPLTYPVKRTTKMAGMMIAAAWVLSFILWAPAILFWQFI--VGVRTVEDGECYIQF---------FSNAAVTFGTAIAAFYLPVIIMTVLYWHISRASKSR----REKKVTRTILAILLAFIITWAPYNVMVLINTFCAPCIPNTVWTIGYWLCYINSTINPACYALCNATFKKTFKHLLM- | |||||||||||||||||||
| 6 | 3emlA | 0.17 | 0.22 | 0.87 | 2.63 | Download | --------------------------MGSSVYITVELAIAVLAILGNVLVCWAVWLNSNLQNVTNYFVVSLAAADIAVGVLAIPFAITIST--GFCAACHGCLFIACFVLVLTQSSIFSLLAIAIDRYIAIRIPLRYNGLVTGTRAKGIIAICWVLSFAIGLTPMLGWNNCGGCGEGQVACLFEDVV-----------PMNYMVYFNFFACVLVPLLLMLGVYLRIFLAARRQLVHAAKSLAIIVGLFALCWLPLHIINCFTFFCPDC---SHAPLWLMYLAIVLSHTNSVVNPFIYAYRIREFRQTFRKIIRS | |||||||||||||||||||
| 7 | 3d4s | 0.20 | 0.22 | 0.62 | 1.70 | Download | ----------------------------VVGMGIVMSLIVLAIVFGNVLVITAIAKFERLQTVTNYFITSLACADLVMGLAVVPFGAAHILMKMWTFGNFWCEFWTSIDVLCVTASIWTLCVIAVDRYFAITSPFKYQSLLTKNKARVIILMVWIVSGLTSFLPIQMWYRAT--H---QEAINCYAEE----TCCDFFTNQAYAIASSIVSFYVPLVIMVFVYSRVFQEAKR---------------------------------------------------------------------------------- | |||||||||||||||||||
| 8 | 4buoA | 0.15 | 0.21 | 0.94 | 2.94 | Download | -------NSDL---------DVNTDIYSKVLVTAIYLALFVVGTVGNSVTLFTLARKKSLQSTVDYYLGSLALSDLLILLLAMPVELYNFIWVHHPWAFAGCRGYYFLRDACTYATALNVVSLSVELYLAICHPFKAKTLMSRSRTKKFISAIWLASALLAIPMLFTMGLQNLSGDGTHPGGLVCTPIVDTATLKVVIQVNTFMSFLFPMLVASILNTVIANKLTVMVHQPGRVQALRRGVLVLRAVVIAFVVCWLPYHVRRLMFCYISDEQWTYHYFYMLTNALVYVSAAINPILYNLVSANFRQVFLSTL-- | |||||||||||||||||||
| 9 | 5tgzA | 0.19 | 0.24 | 0.91 | 3.16 | Download | --GGRGENFMDIEC--FMVLN----PSQQLAIAVLSLTLGTFTVLENLLVLCVILHSRSLRRPSYHFIGSLAVADLLGSVIFVYSFIDFHVFHRK-DSRNVFLFKLGGVTASFTASVGSLFLAAIDRYISIHRPLAYKRIVTRPKAVVAFCLMWTIAIVIAVLPLLG--------NCEKLQSVC---------SDIFPHIDKTYLMFWIGVVSVLLLFIVYAYMYILWKA-APDQARMDIELAKTLVLILVVLIICWGPLLAIMVYDVFGKMNKIKTVFAFCSMLCLLNSTVNPIIYALRSKDLRHAFRSMF-- | |||||||||||||||||||
| 10 | 4gpoA | 0.19 | 0.19 | 0.89 | 4.73 | Download | ------------------------LQQWEAGMSLLMALVVLLIVAGNVLVIAAIGSTQRLQTLTNLFITSLACADLVVGLLVVPFGATLVVRGTWLWGSFLCELWTSLDVLCVTASIETLCVIAIDRYLAITSPFRYQSLMTRARAKVIICTVWAISALVSFLPIMMHWWRDEDPQALKCYQDPG--------CCDFVTNRAYAIASSIISFYIPLLIMIFVALRVYREAKEQVMLMREHKALKTLGIIMGVFTLCWLPFFLVNIVNVFNRDLVPDWLFVAFNWLGYANSAMNPIIYC-RSPDFRKAFKRL--- | |||||||||||||||||||
| ||||||||||||||||||||||||||
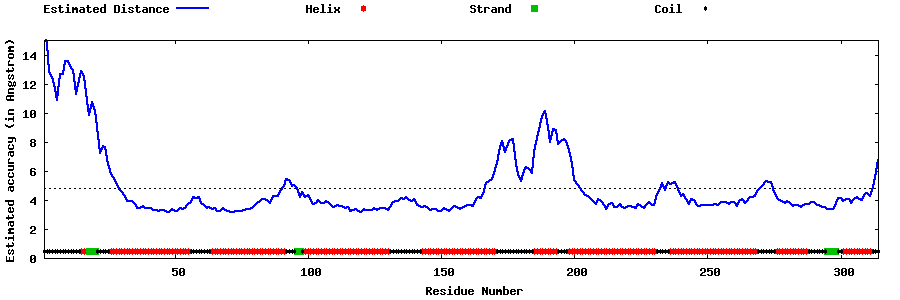
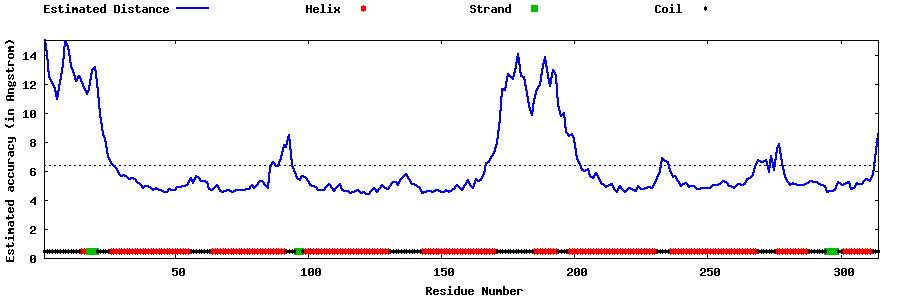
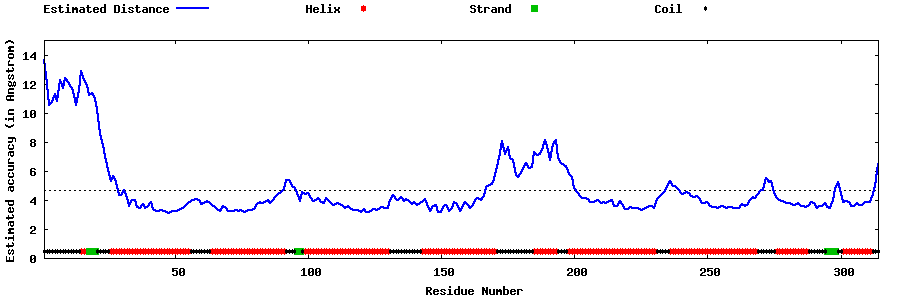
| Generated 3D models | Estimated local accuracy of models | ||
 | |||
|
|
|||
 | |||
|
| |||
 | |||
|
| |||
 | |||
|
| |||
 | |||
|
| |||