

| 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 320 340 | |
| | | | | | | | | | | | | | | | | | | |
| MDSTFTGYNLYNLQVKTEMDKLSSGLDIYRNPLKNKTEVTMFILTGFTDDFELQVFLFLLFFAIYLFTLIGNLGLVVLVIEDSWLHNPMYYFLSVLSFLDACYSTVVTPKMLVNFLAKNKSISFIGCATQMLLFVTFGTTECFLLAAMAYDHYVAIYNPLLYSVSMSPRVYVPLITASYVAGILHATIHIVATFSLSFCGSNEIRHVFCDMPPLLAISCSDTHTNQLLLFYFVGSIEIVTILIVLISCDFILLSILKMHSAKGRQKAFSTCGSHLTGVTIYHGTILVSYMRPSSSYASDHDIIVSIFYTIVIPKLNPIIYSLRNKEVKKAVKKMLKLVYK | |
| CCCCCCCCCHHHHHHHHHHHHHCCCCCHHHCCCCCCCCCCSSSSSCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCHHHHHHHHHHHHCCCCCCCCHHHHHHHHCCCCSSCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHSSCCCCCCCCCCCCCCHHHHHHHCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHSSCCCCCCCCCHHHCHHHHHHHHHHHHHHHSSSSSCCCCCCCCCCCCSSSSSSCCHHHCCCCHHHCCCCHHHHHHHHHHHHHHCC | |
| 9743135312425665300100143336442477884120346865889936089999999999999999887999998508886674888877799884472125718898987148978848999999999999999999999999864134106210288026788999999999999999999999998453089999138541581888877047862767889999999999999999999999999981266747664432155889997999873221458378999878789889988220031243574505649899999999998659 |
| 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 320 340 | |
| | | | | | | | | | | | | | | | | | | |
| MDSTFTGYNLYNLQVKTEMDKLSSGLDIYRNPLKNKTEVTMFILTGFTDDFELQVFLFLLFFAIYLFTLIGNLGLVVLVIEDSWLHNPMYYFLSVLSFLDACYSTVVTPKMLVNFLAKNKSISFIGCATQMLLFVTFGTTECFLLAAMAYDHYVAIYNPLLYSVSMSPRVYVPLITASYVAGILHATIHIVATFSLSFCGSNEIRHVFCDMPPLLAISCSDTHTNQLLLFYFVGSIEIVTILIVLISCDFILLSILKMHSAKGRQKAFSTCGSHLTGVTIYHGTILVSYMRPSSSYASDHDIIVSIFYTIVIPKLNPIIYSLRNKEVKKAVKKMLKLVYK | |
| 6432333341231313330441375243440345030200100000004324010000130332132133233100100100030000000001000000010000000300000015432010200010010000000000000010001000000200102040033000000000001001001010000030201431201000003100010001213010100011013223302300330321001000103026323100000000210010231012000000214303444100001102100230020000114301400330043318 |
| Rank | PDB Hit | Iden1 | Iden2 | Cov. | Norm. Z-score | Download Align. | 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 320 340 | | | | | | | | | | | | | | | | | | |||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sec.Str Seq | CCCCCCCCCHHHHHHHHHHHHHCCCCCHHHCCCCCCCCCCSSSSSCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCHHHHHHHHHHHHCCCCCCCCHHHHHHHHCCCCSSCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHSSCCCCCCCCCCCCCCHHHHHHHCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHSSCCCCCCCCCHHHCHHHHHHHHHHHHHHHSSSSSCCCCCCCCCCCCSSSSSSCCHHHCCCCHHHCCCCHHHHHHHHHHHHHHCC MDSTFTGYNLYNLQVKTEMDKLSSGLDIYRNPLKNKTEVTMFILTGFTDDFELQVFLFLLFFAIYLFTLIGNLGLVVLVIEDSWLHNPMYYFLSVLSFLDACYSTVVTPKMLVNFLAKNKSISFIGCATQMLLFVTFGTTECFLLAAMAYDHYVAIYNPLLYSVSMSPRVYVPLITASYVAGILHATIHIVATFSLSFCGSNEIRHVFCDMPPLLAISCSDTHTNQLLLFYFVGSIEIVTILIVLISCDFILLSILKMHSAKGRQKAFSTCGSHLTGVTIYHGTILVSYMRPSSSYASDHDIIVSIFYTIVIPKLNPIIYSLRNKEVKKAVKKMLKLVYK | |||||||||||||||||||||||||
| 1 | 3emlA | 0.19 | 0.19 | 0.81 | 3.26 | Download | --------------------------------------------------IMGSSVYITVELAIAVLAILGNVLVCWAVWLNSNLQNVTNYFVVSLAAADIAVGVLAIPFAITIS---GFCAACHGCLFIACFVLVLTQSSIFSLLAIAIDRYIAIRIPLRYNGLVTGTRAKGIIAICWVLSFAIGLTPMLGWNNCGGCGEGQVACLFEDVVP-----------MNYMVYFNFFACVLVPLLLMLGVYLRIFLAARRQLVHAAKSLAI-IVGLFALCWLPLHIINCFTFFCPDCSHAPLWLMYLAIVLSHTNSVVNPFIYAYRIREFRQTFRKIIRSHVL | |||||||||||||||||||
| 2 | 5tgzA | 0.20 | 0.22 | 0.81 | 1.88 | Download | -------------------------------RGENFMDIECF----MVLNPSQQLAIAVLSLTLGTFTVLENLLVLCVILHSRSLRCPSYHFIGSLAVADLLGSVIFVYSFIDFHVFHRK-DSRNVFLFKLGGVTASFTASVGSLFLAAIDRYISIHRPLAYKRIVTRPKAVVAFCLMWTIAIVIA------------------------VLPLLGWNCEKLQSVDKTYLMFWIGVVSVLLLFIVYAYMYILWKAHSHAPDQARMELAKTLVLILVVLIICWGPLLAIMVYDVFGKMNKLIKTVFAMLCLLNSTVNPIIYALRSKDLRHAFRSM------ | |||||||||||||||||||
| 3 | 5tgzA | 0.18 | 0.22 | 0.82 | 2.09 | Download | -----------------------------GGRGENFMD---------IENPSQQLAIAVLSLTLGTFTVLENLLVLCVILHSRSLRCPSYHFIGSLAVADLLGSVIFVYSFIDFH-VFHRKDSRNVFLFKLGGVTASFTASVGSLFLAAIDRYISIHRPLAYKRIVTRPKAVVAFCLMWTIAIVIAVLPLLG----WNCEKLQSV---C---------SDIFPHIDKTYLMFWIGVVSVLLLFIVYAYMYILWKAHSHAQARMDIELAKTLVLILVVLIICWGPLLAIMVYDVFGKMNKTVFAFCSMLCLLNSTVNPIIYALRSKDLRHAFRSMF----- | |||||||||||||||||||
| 4 | 4djh | 0.13 | 0.18 | 0.82 | 1.53 | Download | -------------------------------------------------SPAIPVIITAVYSVVFVVGLVGNSLVMFVIIRYTKMKTATNIYIFNLALADALVTT-TMPFQSTVYLMNSWPFGDVLCKIVLSIDYYNMFTSIFTLTMMSVDRYIAVCHPVKALDFRTPLKAKIINICIWLLSSSVGISAIVLGGTKVR---EDVDVIECSLQFPD----DDYWWDLFMKICVFIFAFVIPVLIIIVCYTLMILRLKSVRLDRNLRRITRLVLVVVAVFVVCWTPIHIFILVEALGSAALSSYYFCIALGYTNSSLNPILYAFLDENFKRCFRDFCFP--- | |||||||||||||||||||
| 5 | 4yay | 0.14 | 0.23 | 0.92 | 1.26 | Download | DKSPDSPEKDFRHGFDILVGNEGKVKEAEQLKTTRNAYIQKYLILNSSDCPYIFVMIPTLYSIIFVVGIFGNSLVVIVIYFYMKLKTVASVFLLNLALADLCFLLTLPLWAVYTAMEYRWPFGNYLCKIASASVSFNLYASVFLLTCLSIDRYLAIVHPMKSRLRRTMLVAKVTCIIIWLLAGLASLPAIIH-RNVFF------IITVCAFHYE--------TLPIGLGLTKNILGFLFPFLIILTSYTLIWKALK------KNDDIFKIIMAIVLFFFFSWIPHQIFTFLDLGIDIVDTAMPITICIAYFNNCLNPLFYGFLGKKFKRYFLQLL----- | |||||||||||||||||||
| 6 | 3emlA | 0.19 | 0.19 | 0.81 | 3.47 | Download | ---------------------------------------------------MGSSVYITVELAIAVLAILGNVLVCWAVWLNSNLQNVTNYFVVSLAAADIAVGVLAIPFAIT--ISTGFCAACHGCLFIACFVLVLTQSSIFSLLAIAIDRYIAIRIPLRYNGLVTGTRAKGIIAICWVLSFAIGLTPMLGWNNCGGCGEGQVACLFEDVVP-----------MNYMVYFNFFACVLVPLLLMLGVYLRIFLAARRQLVHAAKSLAIIVGLFALCWLPLHIINCF-TFFCPDCSHAPLWLMYLAIVLSHTNSVVNPFIYAYRIREFRQTFRKIIRSHVL | |||||||||||||||||||
| 7 | 4iaq | 0.21 | 0.21 | 0.77 | 1.72 | Download | ---------------------------------------------------PWKVLLVMLLALITLATTLSNAFVIATVYRTRKLHTPANYLIASLAVTDLLVSILVMPISTMYTVTGRWTLGQVVCDFWLSSDITCCTASIWHLCVIALDRYWAITDAVEYSAKRTPKRAAVMIALVWVFSISISLPPFF-WRQASECVVNT--------------------DHILYTVYSTVGAFYFPTLLLIALYGRIYVEARSRIAAARERKATKTLGIILGAFIVCWLPFFIISLVMPIH--LAIF-DFFTWLGYLNSLINPIIYTMSNEDFKQAFHKLIRF--- | |||||||||||||||||||
| 8 | 2z73A | 0.18 | 0.21 | 0.86 | 2.75 | Download | ---------------------------------ENPSIVVHPHWREFDQVPAVYYSLGIFIGICGIIGCGGNGIVIYLFTKTKSLQTPANMFIINLAFSDFTFSLVNGPLMTISCFLKKWIFGFAACKVYGFIGGIFGFMSIMTMAMISIDRYNVIGRPMAASKKMSHRRAFIMIIFVWLWSVLWAIGPIFGWGAYTLEGVL----------------CSRDSTTRSNILCMFILGFFGPILIIFFCYFNIVMAQAGANAEMRLAKISIVIVSQFLLSWSPYAVVALLAQFGPLEWVTPYAAQLPVMFAKASAIHNPMIYSVSHPKFREAISQTFPWVLT | |||||||||||||||||||
| 9 | 3emlA | 0.19 | 0.19 | 0.80 | 4.77 | Download | ----------------------------------------------------ISSVYITVELAIAVLAILGNVLVCWAVWLNSNLQNVTNYFVVSLAAADIAVGVLAIPFAITIS---GFCAACHGCLFIACFVLVLTQSSIFSLLAIAIDRYIAIRIPLRYNGLVTGTRAKGIIAICWVLSFAIGLTPMLNCGQSQGCGEGQVACLFEDVVP-----------MNYMVYFNFFACVLVPLLLMLGVYLRIFLAARRQLVHAAKSLAII-VGLFALCWLPLHIINCFTFFCPDCSHAPLWLMYLAIVLSHTNSVVNPFIYAYRIREFRQTFRKIIRSHVL | |||||||||||||||||||
| 10 | 2ydoA | 0.16 | 0.19 | 0.84 | 5.34 | Download | -----------------------------------------------------SSVYITVELAIAVLAILGNVLVCWAVWLNSNLQNVTNYFVVSAAAADILVGVLAIPFAIA--ISTGFCAACHGCLFIACFVLVLTASSIFSLLAIAIDRYIAIRIPLRYNGLVTGTRAKGIIAICWVLSFAIGLTPMLGWNNCGQPKEGKAHSQGCGEGQVACLFEDVVPMNYMVYFNFFACVLVPLLLMLGVYLRIFLAARRQLKQMESTLQKEVHAAKSLAIIVGLFALCWFTFFCPDCSHAPLWLMYLAIVLSHTNSVVNPFIYAYRIREFRQTFRKIIRSHVL | |||||||||||||||||||
| ||||||||||||||||||||||||||
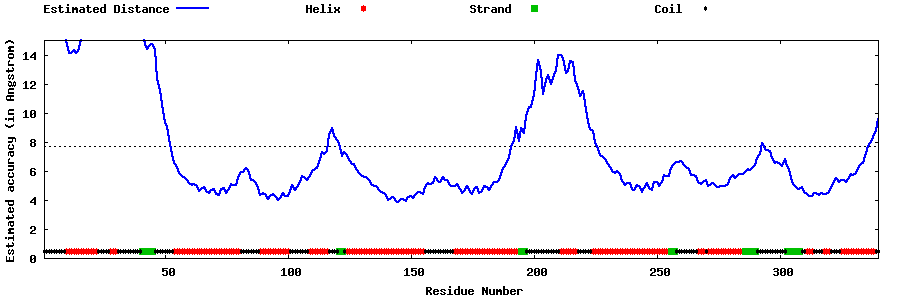
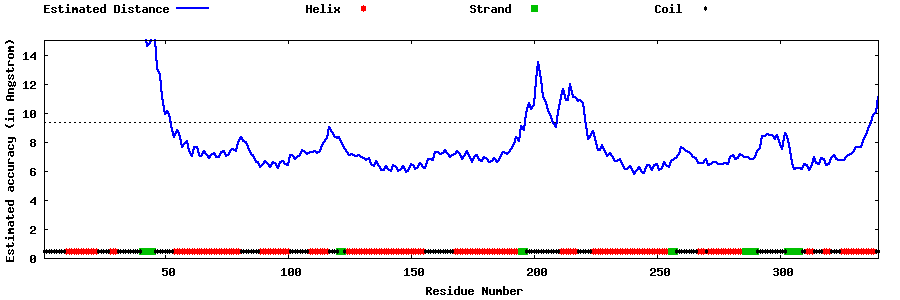
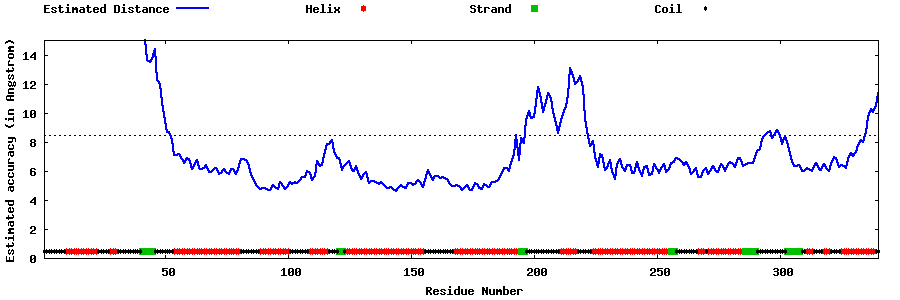
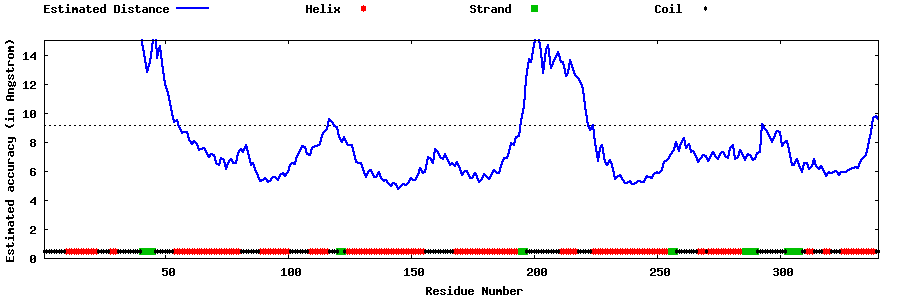
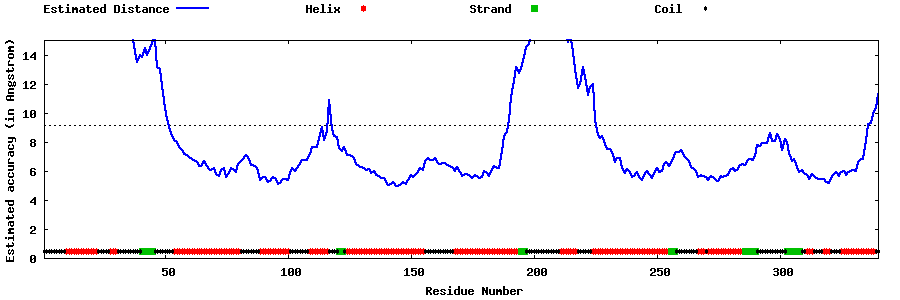
| Generated 3D models | Estimated local accuracy of models | ||
 | |||
|
|
|||
 | |||
|
| |||
 | |||
|
| |||
 | |||
|
| |||
 | |||
|
| |||