

| 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 | |
| | | | | | | | | | | | | | | | | |
| MDGDNQSENSQFLLLGISESPEQQQILFWMFLSMYLVTVLGNVLIILAISSDSHLHTPMYFFLANLSFTDLFFVTNTIPKMLVNFQSQNKAISYAGCLTQLYFLVSLVTLDNLILAVMAYDRYVAICCPLHYVTAMSPGLCVLLLSLCWGLSVLYGLLLTFLLTRVTFCGPREIHYLFCDMYILLWLACSNTHIIHTALIATGCFIFLTLLGFMTTSYVRIVRTILQMPSASKKYKTFSTCASHLGVVSLFYGTLAMVYLQPLHTYSMKDSVATVMYAVLTPMMNPFIYSLRNKDMHGAPGRVLWRPFQRP | |
| CCCCCCCCCHHHHSSCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCHHHHHHHHHHHHHCHHCCCHHHHHHHHHCCCCSSCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHSSCCCCCCCSSCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCSSCCCCCCHHHHHHHHCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHSSCCCCCCCCCSSSCCHHHHHHHHHHHCCHHSSSSCCCCCCCCCCCSSSSSSCCCCCCCCCHHHCCCCHHHHHHHHHHHCCCCCCC | |
| 98988851644520179988427789999999999999999999999997178755629999888998754514157899889983589667388899999999999999999999998751432376543785237618999999999999999999999997416899894488267868998671545379999999999999999899999999999988311467788675205716688889998571432670789778888847998501536254455414661889999999970655899 |
| 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 | |
| | | | | | | | | | | | | | | | | |
| MDGDNQSENSQFLLLGISESPEQQQILFWMFLSMYLVTVLGNVLIILAISSDSHLHTPMYFFLANLSFTDLFFVTNTIPKMLVNFQSQNKAISYAGCLTQLYFLVSLVTLDNLILAVMAYDRYVAICCPLHYVTAMSPGLCVLLLSLCWGLSVLYGLLLTFLLTRVTFCGPREIHYLFCDMYILLWLACSNTHIIHTALIATGCFIFLTLLGFMTTSYVRIVRTILQMPSASKKYKTFSTCASHLGVVSLFYGTLAMVYLQPLHTYSMKDSVATVMYAVLTPMMNPFIYSLRNKDMHGAPGRVLWRPFQRP | |
| 86572313002000010144262120000100301220233120000001001200000000000101101112003203002012143340302000001100110022010000000310000001002000000030000001200210020011000002000103724000000123200300011021010001221333133233003201010000001021462321000000000000001110000000000223343220000000231233001000020310230023004333748 |
| Rank | PDB Hit | Iden1 | Iden2 | Cov. | Norm. Z-score | Download Align. | 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 | | | | | | | | | | | | | | | | |||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sec.Str Seq | CCCCCCCCCHHHHSSCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCHHHHHHHHHHHHHCHHCCCHHHHHHHHHCCCCSSCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHSSCCCCCCCSSCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCSSCCCCCCHHHHHHHHCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHSSCCCCCCCCCSSSCCHHHHHHHHHHHCCHHSSSSCCCCCCCCCCCSSSSSSCCCCCCCCCHHHCCCCHHHHHHHHHHHCCCCCCC MDGDNQSENSQFLLLGISESPEQQQILFWMFLSMYLVTVLGNVLIILAISSDSHLHTPMYFFLANLSFTDLFFVTNTIPKMLVNFQSQNKAISYAGCLTQLYFLVSLVTLDNLILAVMAYDRYVAICCPLHYVTAMSPGLCVLLLSLCWGLSVLYGLLLTFLLTRVTFCGPREIHYLFCDMYILLWLACSNTHIIHTALIATGCFIFLTLLGFMTTSYVRIVRTILQMPSASKKYKTFSTCASHLGVVSLFYGTLAMVYLQPLHTYSMKDSVATVMYAVLTPMMNPFIYSLRNKDMHGAPGRVLWRPFQRP | |||||||||||||||||||||||||
| 1 | 3emlA | 0.21 | 0.23 | 0.89 | 3.62 | Download | --------------------IMGSSVYITVELAIAVLAILGNVLVCWAVWLNSNLQNVTNYFVVSLAAADIAVGVLAIPFAITIST--GFCAACHGCLFIACFVLVLTQSSIFSLLAIAIDRYIAIRIPLRYNGLVTGTRAKGIIAICWVLSFAIGLTPMLGWNNCGGCGEGQVACLFEDVVP-----------MNYMVYFNFFACVLVPLLLMLGVYLRIFLAARRQLVHAAKSLAIIVGLFALCWLPLHIINCFT-FFCPDCSHAPWLMYLAIVLSHTNSVVNPFIYAYRIREFRQTFRKIIRSHVLRQ | |||||||||||||||||||
| 2 | 5tgzA | 0.22 | 0.23 | 0.90 | 2.12 | Download | -RGENFMDIECF----MVLNPSQQLAIAVLSLTLGTFTVLENLLVLCVILHSRSLRCPSYHFIGSLAVADLLGSVIFVYSFIDF-HVFHRKDSRNVFLFKLGGVTASFTASVGSLFLAAIDRYISIHRPLAYKRIVTRPKAVVAFCLMWTIAIVIAVLPLL-----------------GWNCEKLQSVCSDIFPHDKTYLMFWIGVVSVLLLFIVYAYMYILWKAHSHDQARMDIELAKTLVLILVVLIICWGPLLAIMVYDVFGKMNKLIKTVFMLCLLNSTVNPIIYALRSKDLRHAFRSMF------- | |||||||||||||||||||
| 3 | 5tgzA | 0.22 | 0.23 | 0.87 | 2.05 | Download | GRGENFMD---------IENPSQQLAIAVLSLTLGTFTVLENLLVLCVILHSRSLRCPSYHFIGSLAVADLLGSVIFVYSFIDFH-VFHRKDSRNVFLFKLGGVTASFTASVGSLFLAAIDRYISIHRPLAYKRIVTRPKAVVAFCLMWTIAIVIAVLPLLG----WNCEKLIFPHID------------------KTYLMFWIGVVSVLLLFIVYAYMYILWKAHSHAQARMDIELAKTLVLILVVLIICWGPLLAIMVYDVFGKMKTVFAFCSMLCLLNSTVNPIIYALRSKDLRHAFRSMF------- | |||||||||||||||||||
| 4 | 4djh | 0.15 | 0.23 | 0.90 | 1.54 | Download | -------------------SPAIPVIITAVYSVVFVVGLVGNSLVMFVIIRYTKMKTATNIYIFNLALADALVTT-TMPFQSTVYLMNSWPFGDVLCKIVLSIDYYNMFTSIFTLTMMSVDRYIAVCHPVKALDFRTPLKAKIINICIWLLSSSVGISAIVLGGTKVRE---DVDVIECSLQFPDD---DYSWWDLFMKICVFIFAFVIPVLIIIVCYTLMILRLKSVRLLRNLRRITRLVLVVVAVFVVCWTPIHIFILVEALGAALSSYYFCIALGYTNSSLNPILYAFLDENFKRCFRDFCFP----- | |||||||||||||||||||
| 5 | 4yay | 0.17 | 0.24 | 0.91 | 1.24 | Download | KTTRN-AYIQKYLILNSSDCPYIFVMIPTLYSIIFVVGIFGNSLVVIVIYFYMKLKTVASVFLLNLALADLCFLLTLPLWAVYTAMEYRWPFGNYLCKIASASVSFNLYASVFLLTCLSIDRYLAIVHPMKSRLRRTMLVAKVTCIIIWLLAGLASLPAIIHRNV-FF------IITVCAFHYE--------TLPIGLGLTKNILGFLFPFLIILTSYTLIWKALK------KNDDIFKIIMAIVLFFFFSWIPHQIFTFLDVLIRIVTAMPITICIAYFNNCLNPLFYGFLGKKFKRYFLQLL------- | |||||||||||||||||||
| 6 | 3emlA | 0.21 | 0.23 | 0.89 | 3.88 | Download | ---------------------MGSSVYITVELAIAVLAILGNVLVCWAVWLNSNLQNVTNYFVVSLAAADIAVGVLAIPFAIT--ISTGFCAACHGCLFIACFVLVLTQSSIFSLLAIAIDRYIAIRIPLRYNGLVTGTRAKGIIAICWVLSFAIGLTPMLGWNNCGGCGEGQVACLFEDVVP-----------MNYMVYFNFFACVLVPLLLMLGVYLRIFLAARRQLVHAAKSLAIIVGLFALCWLPLHIINCF-TFFCPDCSHAPWLMYLAIVLSHTNSVVNPFIYAYRIREFRQTFRKIIRSHVLRQ | |||||||||||||||||||
| 7 | 4iaq | 0.20 | 0.23 | 0.85 | 1.73 | Download | ---------------------PWKVLLVMLLALITLATTLSNAFVIATVYRTRKLHTPANYLIASLAVTDLLVSILVMPISTMYTVTGRWTLGQVVCDFWLSSDITCCTASIWHLCVIALDRYWAITDAVEYSAKRTPKRAAVMIALVWVFSISISLPPFFW-RQASE----------CVVNT-------DHI---LYTVYSTVGAFYFPTLLLIALYGRIYVEARSRIADARERKATKTLGIILGAFIVCWLPFFIISLVMPIH-LAIFD-FFTWLGYLNSLINPIIYTMSNEDFKQAFHKLIRFK---- | |||||||||||||||||||
| 8 | 2z73A | 0.19 | 0.23 | 0.94 | 2.94 | Download | -ETYNPSIVVHPHWREFDQPDAVYYSLGIFIGICGIIGCGGNGIVIYLFTKTKSLQTPANMFIINLAFSDFTFSLVNGFPLMTISCFLKKIFGFAACKVYGFIGGIFGFMSIMTMAMISIDRYNVIGRPMAASKKMSHRRAFIMIIFVWLWSVLWAIGPIFG------WGAYTLEGVLCD---------STTRSNILCMFILG---FFGPILIIFFCYFNIVMAQAGANAEMRLAKISIVIVSQFLLSWSPYAVVALLAFGPLEWVTPYAAQLPVMFAKASAIHNPMIYSVSHPKFREAISQTFPWVLTCC | |||||||||||||||||||
| 9 | 3emlA | 0.21 | 0.23 | 0.89 | 4.99 | Download | -------------IM-------GSSVYITVELAIAVLAILGNVLVCWAVWLNSNLQNVTNYFVVSLAAADIAVGVLAIPFAITIST--GFCAACHGCLFIACFVLVLTQSSIFSLLAIAIDRYIAIRIPLRYNGLVTGTRAKGIIAICWVLSFAIGLTPMLNCGQSQGCGEGQVACLFEDVVP-----------MNYMVYFNFFACVLVPLLLMLGVYLRIFLAARRQLVHAAKSLAIIVGLFALCWLPLHI-INCFTFFCPDCSHAPL-MYLAIVLSHTNSVVNPFIYAYRIREFRQTFRKIIRSHVLRQ | |||||||||||||||||||
| 10 | 2ydoA | 0.18 | 0.23 | 0.92 | 5.73 | Download | -----------------S------SVYITVELAIAVLAILGNVLVCWAVWLNSNLQNVTNYFVVSAAAADILVGVLAIPFAIA--ISTGFCAACHGCLFIACFVLVLTASSIFSLLAIAIDRYIAIRIPLRYNGLVTGTRAKGIIAICWVLSFAIGLTPMLGWNNCGQPKEGKAHSQGCGEGQVACLFEDVVPMNYMVYFNFFACVLVPLLLMLGVYLRIFLAARRQLKQMESTLQKEVHAAKSLAIIVGLFAINCFTFFCPDCSHAPWLMYLAIVLSHTNSVVNPFIYAYRIREFRQTFRKIIRSHVLRQ | |||||||||||||||||||
| ||||||||||||||||||||||||||
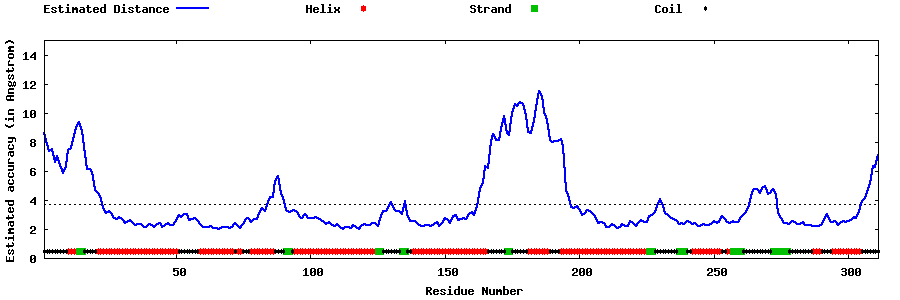
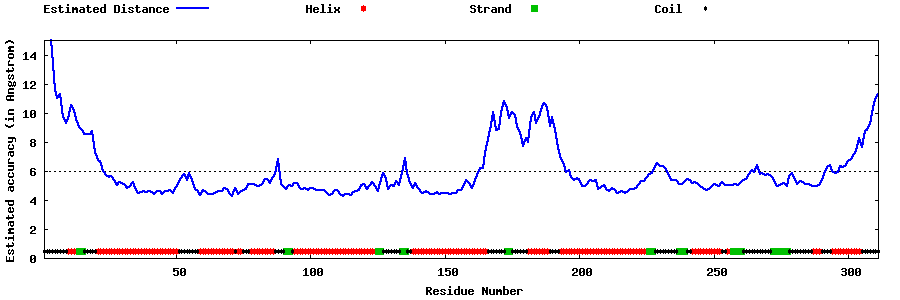
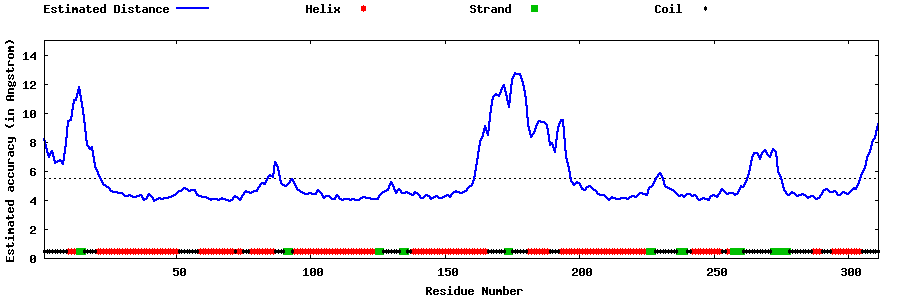
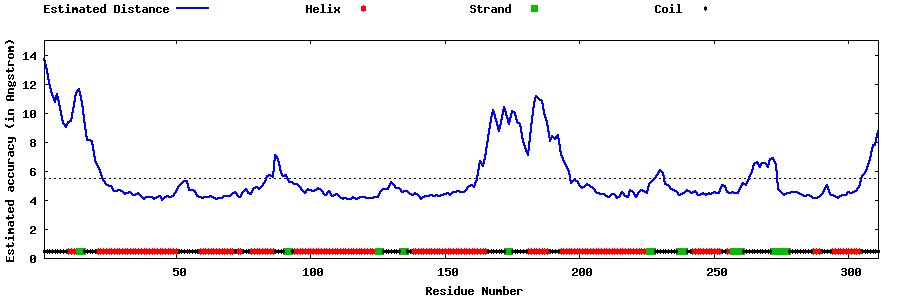
| Generated 3D models | Estimated local accuracy of models | ||
 | |||
|
|
|||
 | |||
|
| |||
 | |||
|
| |||
 | |||
|
| |||
 | |||
|
| |||