

| 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 320 340 360 | |
| | | | | | | | | | | | | | | | | | | | |
| MEDFNMESDSFEDFWKGEDLSNYSYSSTLPPFLLDAAPCEPESLEINKYFVVIIYALVFLLSLLGNSLVMLVILYSRVGRSVTDVYLLNLALADLLFALTLPIWAASKVNGWIFGTFLCKVVSLLKEVNFYSGILLLACISVDRYLAIVHATRTLTQKRYLVKFICLSIWGLSLLLALPVLLFRRTVYSSNVSPACYEDMGNNTANWRMLLRILPQSFGFIVPLLIMLFCYGFTLRTLFKAHMGQKHRAMRVIFAVVLIFLLCWLPYNLVLLADTLMRTQVIQETCERRNHIDRALDATEILGILHSCLNPLIYAFIGQKFRHGLLKILAIHGLISKDSLPKDSRPSFVGSSSGHTSTTL | |
| CCCCCCCCCCCHHCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHCCSSSSCCCCCCCCCCCSSSSSSHHHHHHHHHHHHHHHHHHSSSSCCCCCSSSSCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCHHHHHHHHHHHHHCCCCCCCCCCCCCCCCCCCCCCCCCCCCC | |
| 987668866530102677777776777888766677777766421778658999999999998779997898875366686889999999999999999689999997379978548899999999999999999999999967044500462235543033364369999999999699997523786999479960389512679999999999999999999999999999999864567665418999999999999997899999999999970678897579999999999999999999979899999438989999999953347787666778899988889999978999 |
| 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 320 340 360 | |
| | | | | | | | | | | | | | | | | | | | |
| MEDFNMESDSFEDFWKGEDLSNYSYSSTLPPFLLDAAPCEPESLEINKYFVVIIYALVFLLSLLGNSLVMLVILYSRVGRSVTDVYLLNLALADLLFALTLPIWAASKVNGWIFGTFLCKVVSLLKEVNFYSGILLLACISVDRYLAIVHATRTLTQKRYLVKFICLSIWGLSLLLALPVLLFRRTVYSSNVSPACYEDMGNNTANWRMLLRILPQSFGFIVPLLIMLFCYGFTLRTLFKAHMGQKHRAMRVIFAVVLIFLLCWLPYNLVLLADTLMRTQVIQETCERRNHIDRALDATEILGILHSCLNPLIYAFIGQKFRHGLLKILAIHGLISKDSLPKDSRPSFVGSSSGHTSTTL | |
| 654342443323312333323213243323333343320444334011000020012002302321220010001133321000000000010011000000000000020011010001300011221111000000000300000000002024223110000000001000000100000020244763210010104543220100010111110123122101200010011023154553300000000000001001220000000100230412353042331021001001000131003100010000440141014003322123463245444444444445444336 |
| Rank | PDB Hit | Iden1 | Iden2 | Cov. | Norm. Z-score | Download Align. | 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 320 340 360 | | | | | | | | | | | | | | | | | | | |||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sec.Str Seq | CCCCCCCCCCCHHCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHCCSSSSCCCCCCCCCCCSSSSSSHHHHHHHHHHHHHHHHHHSSSSCCCCCSSSSCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCHHHHHHHHHHHHHCCCCCCCCCCCCCCCCCCCCCCCCCCCCC MEDFNMESDSFEDFWKGEDLSNYSYSSTLPPFLLDAAPCEPESLEINKYFVVIIYALVFLLSLLGNSLVMLVILYSRVGRSVTDVYLLNLALADLLFALTLPIWAASKVNGWIFGTFLCKVVSLLKEVNFYSGILLLACISVDRYLAIVHATRTLTQKRYLVKFICLSIWGLSLLLALPVLLFRRTVYSSNVSPACYEDMGNNTANWRMLLRILPQSFGFIVPLLIMLFCYGFTLRTLFKAHMGQKHRAMRVIFAVVLIFLLCWLPYNLVLLADTLMRTQVIQETCERRNHIDRALDATEILGILHSCLNPLIYAFIGQKFRHGLLKILAIHGLISKDSLPKDSRPSFVGSSSGHTSTTL | |||||||||||||||||||||||||
| 1 | 4mbsA | 0.33 | 0.29 | 0.81 | 3.21 | Download | ------------------------------------PCQKINVKQIAARLLPPLYSLVFIFGFVGNMLVILILINYKRLKSMTDIYLLNLAISDLFFLLTVPFWAHYAAAQWDFGNTMCQLLTGLYFIGFFSGIFFIILLTIDRYLAVVHAVFALKARTVFGVVTSVITWVVAVFASLPNIIFTRSQKEGL-HYTCSSHFPYSQQFWKNFQTLKIVILGLVLPLLVMVICYSGILKTLLRMKEKKRHRDVRLIFTIMIVYFLFWAPYNIVLLLNTFQEFFGL-NNCSSSNRLDQAMQVTETLGMTHCCINPIIYAFVGEEFRNYLLVFFQ------------------------------ | |||||||||||||||||||
| 2 | 4mbsA | 0.33 | 0.29 | 0.81 | 4.09 | Download | -------------------------------------CQKINVKQIAARLLPPLYSLVFIFGFVGNMLVILILINYKRLKSMTDIYLLNLAISDLFFLLTVPFWAHYAAAQWDFGNTMCQLLTGLYFIGFFSGIFFIILLTIDRYLAVVHAVFALKARTVTGVVTSVITWVVAVFASLPNIIFTRSQKEGL-HYTCSSHFPSQYQFWKNFQTLKIVILGLVLPLLVMVICYSGILKTLLRMEEKKRHRDVRLIFTIMIVYFLFWAPYNIVLLLNTFQEFFGLN-NCSSSNRLDQAMQVTETLGMTHCCINPIIYAFVGEEFRNYLLVFFQ------------------------------ | |||||||||||||||||||
| 3 | 4mbsA | 0.33 | 0.29 | 0.81 | 3.79 | Download | ------------------------------------PCQKINVKQIAARLLPPLYSLVFIFGFVGNMLVILILINYKRLKSMTDIYLLNLAISDLFFLLTVPFWAHYAAAQWDFGNTMCQLLTGLYFIGFFSGIFFIILLTIDRYLAVVHAVFALKARTVFGVVTSVITWVVAVFASLPNIIFTRSQKEGL-HYTCSSHFPYQYQFWKNFQTLKIVILGLVLPLLVMVICYSGILKTLLRMKEKKRHRDVRLIFTIMIVYFLFWAPYNIVLLLNTFQEFFGLNN-CSSSNRLDQAMQVTETLGMTHCCINPIIYAFVGEEFRNYLLVFFQ------------------------------ | |||||||||||||||||||
| 4 | 2lnl | 0.85 | 0.70 | 0.82 | 1.60 | Download | -------------------------------------PCMLETETLNKYVVIIAYALVFLLSLLGNSLVMLVILYSRVGRSVTDVYLLNLALADLLFALTLPIWAASKVNGWIFGTFLCKVVSLLKEVNFYSGILLLACISVDRYLAIVHATRTLTQKRHLVKFVCLGCWGLSMNLSLPFFLFRQAYHPNNSSPVCYEVLGNDTAKWRMVLRILPHTFGFIVPLFVMLFCYGFTLRTLFKAHMGQKHRAMRVIFAVVLIFLLCWLPYNLVLLADTLMRTQVIQESCERRNNIGRALDATEILGFLHSCLNPIIYAFIGQNFRHGFLKILAMHG--------------------------- | |||||||||||||||||||
| 5 | 5t1a | 0.34 | 0.29 | 0.78 | 1.17 | Download | ------------------------------------------VKQIGAQLLPPLYSLVFIFGFVGNMLVVLILINCKKLKCLTDIYLLNLAISDLLFLITLPLWAHSAANEWVFGNAMCKLFTGLYHIGYFGGIFFIILLTIDRYLAIVHAVFALKARTTFGVVTSVITWLVAVFASVPGIIFTKQKE--DSVYVCGPYFPRG---WNNFHTIMRNILGLVLPLLIMVICYSGISRASKSRIPSREKKAVRVIFTIMIVYFLFWTPYNIVILLNTFQEFFG-LSNCESTSQLDQATQVTETLGMTHCCINPIIYAFVGEKFRRYLSVFF------------------------------- | |||||||||||||||||||
| 6 | 4mbsA | 0.33 | 0.29 | 0.81 | 3.28 | Download | -------------------------------------CQKINVKQIAARLLPPLYSLVFIFGFVGNMLVILILINYKRLKSMTDIYLLNLAISDLFFLLTVPFWAHYAAAQWDFGNTMCQLLTGLYFIGFFSGIFFIILLTIDRYLAVVHAVFALKARTTFGVVTSVITWVVAVFASLPNIIFTRSQKE-GLHYTCSSHFPSQYQFWKNFQTLKIVILGLVLPLLVMVICYSGILKTLLRMKEKKRHRDVRLIFTIMIVYFLFWAPYNIVLLLNTFQEFFGLN-NCSSSNRLDQAMQVTETLGMTHCCINPIIYAFVGEEFRNYLLVFFQ------------------------------ | |||||||||||||||||||
| 7 | 2lnl | 0.85 | 0.70 | 0.82 | 1.67 | Download | -------------------------------------PCMLETETLNKYVVIIAYALVFLLSLLGNSLVMLVILYSRVGRSVTDVYLLNLALADLLFALTLPIWAASKVNGWIFGTFLCKVVSLLKEVNFYSGILLLACISVDRYLAIVHATRTLTQKRHLVKFVCLGCWGLSMNLSLPFFLFRQAYHPNNSSPVCYEVLGNDTAKWRMVLRILPHTFGFIVPLFVMLFCYGFTLRTLFKAHMGQKHRAMRVIFAVVLIFLLCWLPYNLVLLADTLMRTQVIQESCERRNNIGRALDATEILGFLHSCLNPIIYAFIGQNFRHGFLKILAMH---------------------------- | |||||||||||||||||||
| 8 | 2lnlA | 0.85 | 0.70 | 0.82 | 5.01 | Download | -------------------------------------PCMLETETLNKYVVIIAYALVFLLSLLGNSLVMLVILYSRVGRSVTDVYLLNLALADLLFALTLPIWAASKVNGWIFGTFLCKVVSLLKEVNFYSGILLLACISVDRYLAIVHATRTLTQKRHLVKFVCLGCWGLSMNLSLPFFLFRQAYHPNNSSPVCYEVLGNDTAKWRMVLRILPHTFGFIVPLFVMLFCYGFTLRTLFKAHMGQKHRAMRVIFAVVLIFLLCWLPYNLVLLADTLMRTQVIQESCERRNNIGRALDATEILGFLHSCLNPIIYAFIGQNFRHGFLKILAMHG--------------------------- | |||||||||||||||||||
| 9 | 4mbsA | 0.34 | 0.29 | 0.81 | 3.51 | Download | ------------------------------------PCQKINVKQIAARLLPPLYSLVFIFGFVGNMLVILILINYKRLKSMTDIYLLNLAISDLFFLLTVPFWAHYAAAQWDFGNTMCQLLTGLYFIGFFSGIFFIILLTIDRYLAVVHAVFALKARTVFGVVTSVITWVVAVFASLPNIIFTRSQKE-GLHYTCSSHFPYSYQFWKNFQTLKIVILGLVLPLLVMVICYSGILKTLLRMKEEEKHRDVRLIFTIMIVYFLFWAPYNIVLLLNTFQEFFGLN-NCSSSNRLDQAMQVTETLGMTHCCINPIIYAFVGEEFRNYLLVFFQ------------------------------ | |||||||||||||||||||
| 10 | 5c1mA | 0.32 | 0.27 | 0.79 | 4.07 | Download | --------------------------------GSHSLPQTGSPSMVTAITIMALYSIVCVVGLFGNFLVMYVIVRYTKMKTATNIYIFNLALADALATSTLPFQSVNYLMGWPFGNILCKIVISIDYYNMFTSIFTLCTMSVDRYIAVCHPVKALDFRTPRAKIVNVCNWILSSAIGLPVMFMATTKYR-QGSIDCTLTFSHPTWYWENLLKICVFIFAFIMPVLIITVCYGLMILRLKSVRMRNLRRITRMVLVVVAVFIVCWTPIHIYVIIKAL-------ITIPETTFQTVSWHFCIALGYTNSCLNPVLYAFLDENFKRCF----------------------------------- | |||||||||||||||||||
| ||||||||||||||||||||||||||
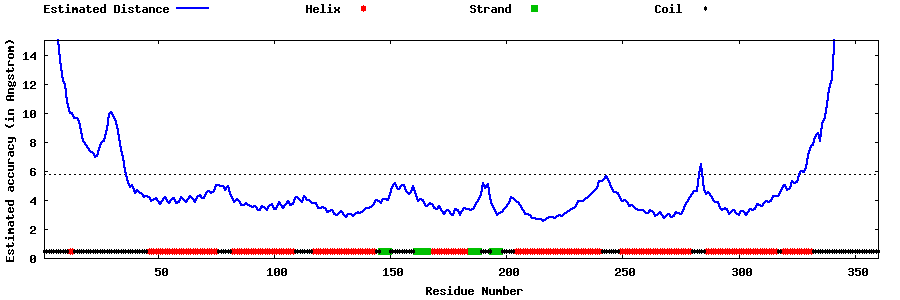
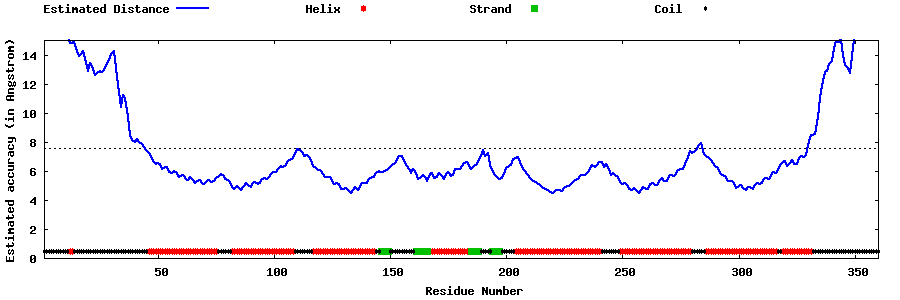
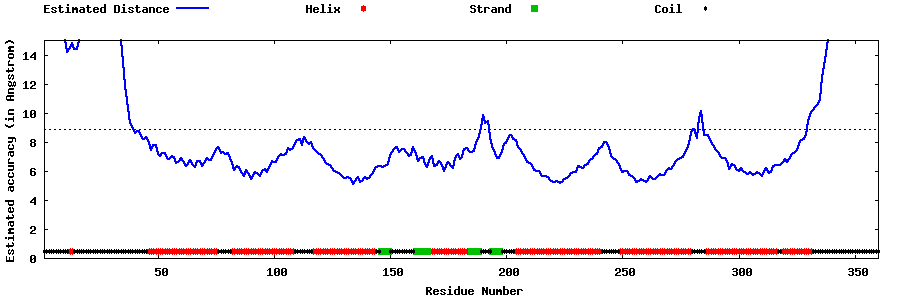
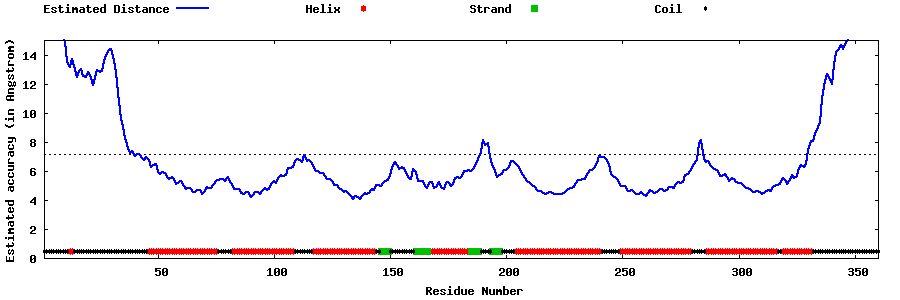
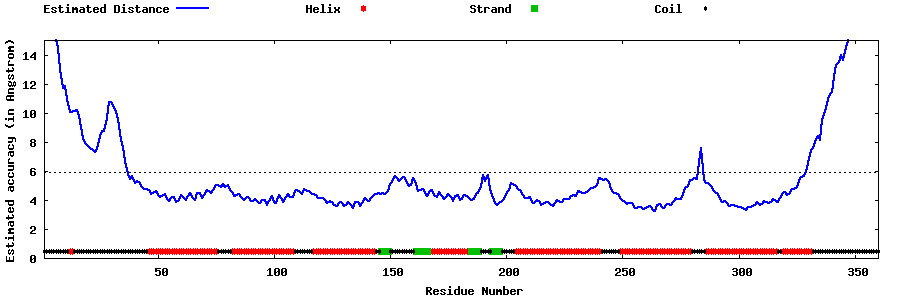
| Generated 3D models | Estimated local accuracy of models | ||
 | |||
|
|
|||
 | |||
|
| |||
 | |||
|
| |||
 | |||
|
| |||
 | |||
|
| |||