

You can:
| Name | 5-hydroxytryptamine receptor 2A |
|---|---|
| Species | Homo sapiens (Human) |
| Gene | HTR2A |
| Synonym | 5-HT-2 serotonin receptor 2A serotonin 5HT-2 receptor 5-HT-2A 5-HT2A receptor [ Show all ] |
| Disease | Depression Unspecified Diabetes Erythropoietic porphyria Fibromyalgia [ Show all ] |
| Length | 471 |
| Amino acid sequence | MDILCEENTSLSSTTNSLMQLNDDTRLYSNDFNSGEANTSDAFNWTVDSENRTNLSCEGCLSPSCLSLLHLQEKNWSALLTAVVIILTIAGNILVIMAVSLEKKLQNATNYFLMSLAIADMLLGFLVMPVSMLTILYGYRWPLPSKLCAVWIYLDVLFSTASIMHLCAISLDRYVAIQNPIHHSRFNSRTKAFLKIIAVWTISVGISMPIPVFGLQDDSKVFKEGSCLLADDNFVLIGSFVSFFIPLTIMVITYFLTIKSLQKEATLCVSDLGTRAKLASFSFLPQSSLSSEKLFQRSIHREPGSYTGRRTMQSISNEQKACKVLGIVFFLFVVMWCPFFITNIMAVICKESCNEDVIGALLNVFVWIGYLSSAVNPLVYTLFNKTYRSAFSRYIQCQYKENKKPLQLILVNTIPALAYKSSQLQMGQKKNSKQDAKTTDNDCSMVALGKQHSEEASKDNSDGVNEKVSCV |
| UniProt | P28223 |
| Protein Data Bank | 6a93, 6a94 |
| GPCR-HGmod model | P28223 |
| 3D structure model | This structure is from PDB ID 6a93. |
| BioLiP | BL0441025,BL0441028, BL0441031, BL0441030,BL0441033, BL0441029,BL0441032, BL0441026, BL0441024,BL0441027 |
| Therapeutic Target Database | T32060 |
| ChEMBL | CHEMBL224 |
| IUPHAR | 6 |
| DrugBank | BE0000451 |
| Name | Ziprasidone |
|---|---|
| Molecular formula | C21H21ClN4OS |
| IUPAC name | 5-[2-[4-(1,2-benzothiazol-3-yl)piperazin-1-yl]ethyl]-6-chloro-1,3-dihydroindol-2-one |
| Molecular weight | 412.936 |
| Hydrogen bond acceptor | 5 |
| Hydrogen bond donor | 1 |
| XlogP | 4.0 |
| Synonyms | CP 88059;CP-88059;CP88059 CPD000466328 DSSTox_CID_3753 146939-27-7 Geodon Oral [ Show all ] |
| Inchi Key | MVWVFYHBGMAFLY-UHFFFAOYSA-N |
| Inchi ID | InChI=1S/C21H21ClN4OS/c22-17-13-18-15(12-20(27)23-18)11-14(17)5-6-25-7-9-26(10-8-25)21-16-3-1-2-4-19(16)28-24-21/h1-4,11,13H,5-10,12H2,(H,23,27) |
| PubChem CID | 60854 |
| ChEMBL | CHEMBL708 |
| IUPHAR | 59 |
| BindingDB | 50048803 |
| DrugBank | DB00246 |
Structure | 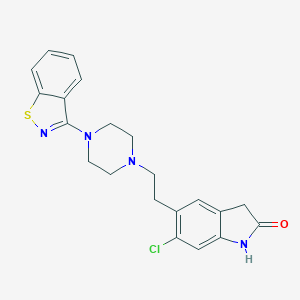 |
| Lipinski's druglikeness | This ligand satisfies Lipinski's rule of five. |
| Parameter | Value | Reference | Database source |
|---|---|---|---|
| N/A | N/A | DrugBank | |
| IC50 | 0.42 nM | PMID16220969 | BindingDB |
| IC50 | 0.42 nM | PMID16220969 | ChEMBL |
| Ki | 0.08 nM | PMID18160289 | PDSP |
| Ki | 0.08 nM | PMID18160289 | BindingDB,ChEMBL |
| Ki | 0.12 nM | PMID11132243 | PDSP,BindingDB |
| Ki | 0.25 nM | PMID11170639 | BindingDB,ChEMBL |
| Ki | 0.28 nM | , None | BindingDB,ChEMBL |
| Ki | 0.3 nM | PMID14998318 | BindingDB |
| Ki | 0.3 nM | PMID14998318, PMID12629531 | PDSP,BindingDB,ChEMBL |
| Ki | 0.309029 - 1.58489 nM | PMID8935801, PMID12629531, PMID12176106, PMID18308814 | IUPHAR |
| Ki | 0.39 nM | PMID18595716 | BindingDB,ChEMBL |
| Ki | 0.39 nM | PMID18595716 | PDSP |
| Ki | 0.4 nM | PMID23919353 | ChEMBL |
| Ki | 0.5 nM | PMID12176106 | PDSP,BindingDB |
| Ki | 0.631 nM | PMID17880057 | ChEMBL |
| Ki | 0.851138 nM | http://www.nature.com/tpj/journal/v6/n1/pdf/6500342a.pdf | PDSP |
| Ki | 1.4 nM | PMID8935801 | PDSP,BindingDB |
zhanglab![]() zhanggroup.org | +65-6601-1241 | Computing 1, 13 Computing Drive, Singapore 117417
zhanggroup.org | +65-6601-1241 | Computing 1, 13 Computing Drive, Singapore 117417